 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_1639_f_10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 451aa MW: 49313 Da PI: 5.3609 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 86.3 | 2.8e-27 | 103 | 159 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+DgynWrKYGqK v+g+ef rsYY+Ct ++Cp+kk++++++ d+++ +++Y geH h+
Neem_1639_f_10 103 EDGYNWRKYGQKLVRGNEFVRSYYKCTNSSCPAKKQLDCTH-DGQIADTIYFGEHCHP 159
7**************************************99.***************7 PP
| |||||||
| 2 | WRKY | 84.6 | 9.5e-27 | 276 | 330 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+++pr C+s+gCp+kk+ver+++dpk+v++tYeg+H+he
Neem_1639_f_10 276 VNDGYRWRKYGQKLVKGNPNPR----CSSSGCPAKKHVERASHDPKLVITTYEGQHDHE 330
58*******************6....********************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 19.703 | 97 | 161 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 6.15E-22 | 97 | 160 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 8.7E-24 | 98 | 160 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.8E-29 | 102 | 160 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.0E-21 | 103 | 159 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 5.0E-29 | 262 | 332 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 2.22E-23 | 268 | 332 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 29.861 | 271 | 332 | IPR003657 | WRKY domain |
| SMART | SM00774 | 2.5E-32 | 276 | 331 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 4.4E-20 | 277 | 329 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 451 aa Download sequence Send to blast |
MVSSPEDVVD KVASDKLQEG PCAESVSETH ALQCDQQVSM PPRIPNKEPM APTDTGHSMP 60 MDQERSISFV TSEKASETTD IVHASQTGQE GSNPMIMREK VSEDGYNWRK YGQKLVRGNE 120 FVRSYYKCTN SSCPAKKQLD CTHDGQIADT IYFGEHCHPR LPNLPLAVGF VVSIVEEKPD 180 DSSSSIAKDK SSDVRGQTPC QIEPTDNPRP SVVSVSDGVK GSPLQSNKIK DEVDNDDRPG 240 SKRRKKDNYN TSATIVERPS GEQRVVVQTL SEIDFVNDGY RWRKYGQKLV KGNPNPRCSS 300 SGCPAKKHVE RASHDPKLVI TTYEGQHDHE MPPSRTVTHN VAGTTSNKTA NNGESATKLE 360 ENDPTCPDTA VHSSSDLPSK SSEEQNSQPG TIEGSDIGQD TVVHASSCHK SGANDKSNCE 420 SETKSEQNGT ACRETNFSEQ EQTPKTEAVQ S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 9e-31 | 101 | 333 | 2 | 75 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
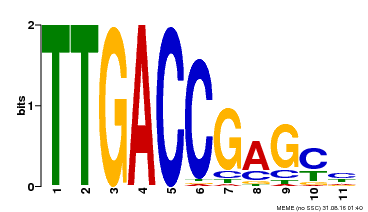
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006447745.1 | 0.0 | WRKY transcription factor 1 | ||||
| Refseq | XP_006469519.1 | 0.0 | WRKY transcription factor 1 | ||||
| Swissprot | Q9SI37 | 2e-78 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | A0A2H5PLC6 | 0.0 | A0A2H5PLC6_CITUN; Uncharacterized protein | ||||
| TrEMBL | V4UA12 | 0.0 | V4UA12_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006469519.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006447745.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7949 | 26 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 8e-78 | zinc-dependent activator protein-1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



