 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_15089_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 387aa MW: 43413.2 Da PI: 9.2086 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 98.6 | 4.3e-31 | 71 | 124 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH +Fv+ave+LGG+e+AtPk +l++m++kgL ++hvkSHLQ+YR
Neem_15089_f_1 71 PRLRWTPELHLCFVHAVERLGGQERATPKLVLQFMDIKGLNISHVKSHLQMYRN 124
8****************************************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-28 | 67 | 125 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.547 | 67 | 127 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.15E-14 | 68 | 125 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.9E-22 | 71 | 126 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.5E-8 | 72 | 123 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 387 aa Download sequence Send to blast |
MKGIDSSERS KTSNSNETGS EDEREEIDKT CQSISGESSS NSSIELENEN KKTPAAGYSG 60 STRQYNRSKT PRLRWTPELH LCFVHAVERL GGQERATPKL VLQFMDIKGL NISHVKSHLQ 120 MYRNKKLDEP NQEQGLLLTH GGGGHQIYNF SQVPTMQNFN RRPSSSLRTL KGSFFHSRCG 180 DVAAWRRYGS QTCSFGLGEN NAVGKAKHGL YASVAERLLG TQNTKLNLHS HTASTCSFNE 240 QDTSKTHQTF EEVDELIRRF RQSHSTSRPS SSMERNFKTQ VEERVKGQIN CSKKNSSHHC 300 NSWPMVQEAE NKLKRKTWDS ECNIDLNLSL KIKPTIDEVL EGAEAQSTSL SLSLSSSSSS 360 KISRLKEGYE SRKQEKTTAA STLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-16 | 72 | 126 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-16 | 72 | 126 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-16 | 72 | 126 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-16 | 72 | 126 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
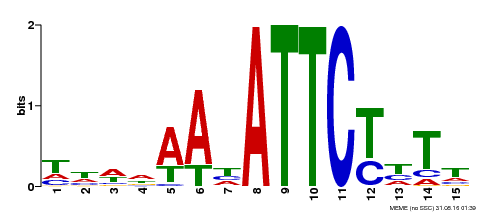
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006486285.1 | 7e-94 | uncharacterized protein LOC102620939 | ||||
| Refseq | XP_024041316.1 | 7e-94 | uncharacterized protein LOC18044186 | ||||
| TrEMBL | A0A2H5NJW4 | 2e-92 | A0A2H5NJW4_CITUN; Uncharacterized protein | ||||
| STRING | XP_006486285.1 | 3e-93 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9359 | 21 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 6e-44 | G2-like family protein | ||||



