 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_11891_f_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 636aa MW: 72669 Da PI: 7.6886 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.8 | 3.8e-28 | 29 | 115 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g++ e+++ ++r++++++t+Cka+++vk++ d+kw ++++ +eHnHel p
Neem_11891_f_3 29 FYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGFTPESDSG---SSRRSSVKKTDCKASMHVKRRADAKWIIHEFIKEHNHELLP 115
9****************************************99888...889999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.8E-25 | 29 | 115 | IPR004330 | FAR1 DNA binding domain |
| PROSITE profile | PS50966 | 9.137 | 352 | 388 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.8E-5 | 363 | 390 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 636 aa Download sequence Send to blast |
MAVVGLEGES SFEPRNGIEF ESHEAAYAFY QEYAKSMGFT TSIKNSRRSK KSKEFIDAKF 60 ACSRYGFTPE SDSGSSRRSS VKKTDCKASM HVKRRADAKW IIHEFIKEHN HELLPALAYH 120 FRIGRNVKLA EKNNIDILHA VSERTRKTYV EMSRQSAGCR NGGFLSDLDN QFDKGRFLPL 180 DEGDAQKCIL KSWTDEEFDM RWWKMVTRFE LQDDEWVQSL YEDRKKWVPT YMGDTFLAGM 240 STTQRSESMN SFFDKYIHKK ITLKEFVKQY GNILQNRYEE EAIADFDTWH KQPALKSPSP 300 WEKQMSTVYT HAIFKKFQVE VLGVVGCHPK KGSEDGTIVT YRVQDCEKNE YFMVTWNELK 360 SEVSCLCRLF ECKGFLCRHA MIVLQICGLS SIPPHYILKR WTKDAKSKQL ISEGAERLQT 420 RVQRYNDLCK RAIELSEEGS LSDENYNIVF RALVDGLKNC VNMNNANNSV LECSSHAQGV 480 REAEETNQGS FATKSSKKKN AIRKRKVQSE PDVLLVDAQE SLQQMENLTS DGITLNGFYA 540 AQQNVQGLVQ LNLMEPPHDG YYVNQQSLQG LVSCAGFMTG QLNSIAPSHD GFFGTQQSMH 600 GLGQLDFRPP TSFGYGLQDE SHLRSTQLHS NAPGHP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
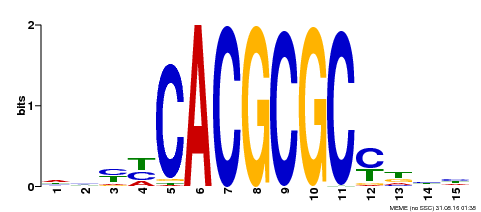
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024037440.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Refseq | XP_024955390.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2P5RM37 | 0.0 | A0A2P5RM37_GOSBA; Uncharacterized protein | ||||
| STRING | VIT_03s0017g01810.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



