 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_021633-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 837aa MW: 90827.8 Da PI: 6.2145 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.7 | 3e-21 | 139 | 194 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+kkl L++rqVk+WFqNrR+++k
AHYPO_021633-RA 139 KKRYHRHTPQQIQELEAVFKECPHPDEKQRLELSKKLCLETRQVKFWFQNRRTQMK 194
688999***********************************************999 PP
| |||||||
| 2 | START | 193 | 1.4e-60 | 349 | 574 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE.....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEE CS
START 1 elaeeaaqelvkkalaeepgWvkss.....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kae 80
ela+ a++elvk+a+++ep+W +s e +n de++++f + + + +ea r++g+v+ ++ lve+++d + +W e+++ +a+
AHYPO_021633-RA 349 ELALGAMDELVKLAQSDEPLWLRSMdsggrEVLNHDEYMRSFNPIIGmkpngFVSEATRETGMVIINSLALVETIMDAN-RWAEMFPgmiaRAT 441
57899************************99***********9977799999***************************.************** PP
EEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEE CS
START 81 tlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwv 167
t++vissg galqlm elq+lsplvp R + f+R+++q+ +g+w++vdvS+d ++ + s+ +v +++lpSg+++++++ng+skvtwv
AHYPO_021633-RA 442 TTDVISSGmggtrnGALQLMHCELQVLSPLVPvRVVNFLRFCKQHAEGVWAVVDVSIDCIRESSISPTYVNCRRLPSGCVVQDMPNGYSKVTWV 535
**************************************************************999***************************** PP
E-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 168 ehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
eh +++++++h+l+r+lv+sgl +ga++w tlqrqce+
AHYPO_021633-RA 536 EHSEYEESSIHQLYRPLVSSGLGFGAQRWITTLQRQCEC 574
*************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.24E-20 | 124 | 196 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.232 | 136 | 196 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-22 | 136 | 204 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.7E-18 | 137 | 200 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.78E-19 | 138 | 196 | No hit | No description |
| Pfam | PF00046 | 9.3E-19 | 139 | 194 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 171 | 194 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.593 | 340 | 577 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.89E-34 | 342 | 574 | No hit | No description |
| CDD | cd08875 | 9.87E-122 | 344 | 573 | No hit | No description |
| SMART | SM00234 | 5.4E-43 | 349 | 574 | IPR002913 | START domain |
| Pfam | PF01852 | 3.9E-52 | 350 | 574 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.43E-24 | 602 | 829 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 837 aa Download sequence Send to blast |
MNFGGFLDSS GGTGSEGGGG GGGARLVTDI PYNNHNMPPG AISPHQQQQQ RLVGPTSMNN 60 KSFFNTPGLS LALQTGMEGH GEVTRVTDQN YEANGNGGGT TIGARRSRDN EEHESRSGSD 120 NLDGASGDEQ DASDLPPRKK RYHRHTPQQI QELEAVFKEC PHPDEKQRLE LSKKLCLETR 180 QVKFWFQNRR TQMKTQLERH ENSILRQEND KLRAENMSIR EALRNPICNN CGGPAMIGDI 240 SMEEQHLRIE NGRLKEELDR VCALASKFIG RPVSSLSGLS APPMPSSSLE LAVGTNGFGP 300 LNTVTTSSLP LAPDFGSGIS NALPVVSPTK STANVSGIER SFERSMYLEL ALGAMDELVK 360 LAQSDEPLWL RSMDSGGREV LNHDEYMRSF NPIIGMKPNG FVSEATRETG MVIINSLALV 420 ETIMDANRWA EMFPGMIARA TTTDVISSGM GGTRNGALQL MHCELQVLSP LVPVRVVNFL 480 RFCKQHAEGV WAVVDVSIDC IRESSISPTY VNCRRLPSGC VVQDMPNGYS KVTWVEHSEY 540 EESSIHQLYR PLVSSGLGFG AQRWITTLQR QCECLAILMS ANIPSGDHNA ITAGGRRSML 600 KLAQRMTNNF CAGVCASTVH KWNKLNMGNI DEDVRVMTRK SMDDPGEPSG IVLSAATSVW 660 LPVTPQRLFD FLRDERLRSE WDILSNGGPM QEMAHIAKGH DQGNCVSLLR AGAVNPNQSS 720 MLILQETCTD ASGSLVVYAP VDIPAMHVVM NGGDSAYVAL LPSGFAIVPD GSGSRGPNIS 780 MDATPQRVGG SLLTVAFQIL VNSLPTAKLT VESVETVSNL ISCTVQKIKA ALNCES* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
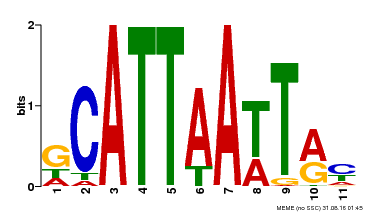
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021849370.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2-like | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A0K9QLI3 | 0.0 | A0A0K9QLI3_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010671720.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_021633-RA |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



