 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_018071-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 540aa MW: 59304.3 Da PI: 6.4252 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 44.4 | 4e-14 | 245 | 304 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +e+Aa+ ++ a++k++g
AHYPO_018071-RA 245 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEEKAARSYDLAALKYWG 304
57*******************666664478446*************************98 PP
| |||||||
| 2 | AP2 | 50.7 | 4.3e-16 | 347 | 398 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.psengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
AHYPO_018071-RA 347 SIYRGVTRHHQQGRWQARIGRvV-G---NKDLYLGTFATEEEAAEAYDIAAIKFRG 398
57****************99944.3...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 6.22E-23 | 245 | 314 | No hit | No description |
| SuperFamily | SSF54171 | 7.19E-17 | 245 | 314 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 5.0E-11 | 245 | 304 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.137 | 246 | 312 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.7E-29 | 246 | 318 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.6E-15 | 246 | 313 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.7E-6 | 247 | 258 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 9.15E-18 | 347 | 407 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.22E-11 | 347 | 406 | No hit | No description |
| Pfam | PF00847 | 6.8E-10 | 347 | 398 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.098 | 348 | 406 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.3E-30 | 348 | 412 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.1E-18 | 348 | 406 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.7E-6 | 388 | 408 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 540 aa Download sequence Send to blast |
MAPAADAANN WLSFSLSPME IYHTPQILPT HPITNNSHNY YLDSFYTPDW GNQKANMGMY 60 TDYLVDQQID NNNPLQTTST VPKLEDFFGG DHLHTTSYNS QTETTTQDSS SQLTHLYESN 120 THQNSVPNYF SNHHTDQYQD LKAMTPSTNT GFQADFSANS GSEVDDSAAT QLGCGANGAE 180 FVGHSIDSTQ TELVPSYATV TDGNNGGSLS LSVVPQSVDK SMVGVGGDSE KCVKKISDTF 240 GQRTSIYRGV TRHRWTGRYE AHLWDNSCRR EGQARKGRQV YLGGYDKEEK AARSYDLAAL 300 KYWGPTATTN FPISNYSNEL EEMKHVTKQE FIASLRRKSS GFSRGASIYR GVTRHHQQGR 360 WQARIGRVVG NKDLYLGTFA TEEEAAEAYD IAAIKFRGVN AVTNFEMKRY DVQAILNSNL 420 PVGGAAKRLK LATEAADQKP TLCLDQQSQS SNGSCGGMSN TINFAAVQSS IPTIASTMPY 480 DTMTAMFHHN LLQHLHSNGG AQAVASDSYG SGVSSMASPT MLLMPQTPAE FYIWPHQSY* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
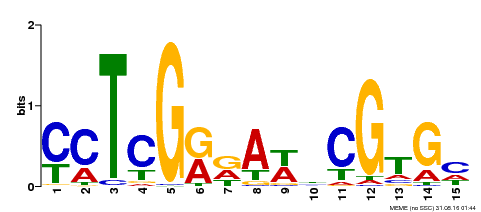
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010680360.2 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 1e-149 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A0R0IAV8 | 0.0 | A0A0R0IAV8_SOYBN; Uncharacterized protein | ||||
| TrEMBL | A0A445J3I4 | 0.0 | A0A445J3I4_GLYSO; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| STRING | XP_010680360.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 1e-144 | AINTEGUMENTA-like 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_018071-RA |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



