 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_017426-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 992aa MW: 111416 Da PI: 6.0734 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.2 | 1.2e-56 | 22 | 136 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
++rwl++ ei++iL n++k+++++e++++p+sgsl+L++rk++ryfrkDG++w+kkkdgkt++E+hekLKv++v++l+cyYah+++n+ fqrr
AHYPO_017426-RA 22 ARHRWLRAAEICEILRNHQKFQISSEPPNQPSSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTIKEAHEKLKVDSVDMLHCYYAHGDDNECFQRR 115
59******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylevk 118
+ywlLeee+ +iv+vhylevk
AHYPO_017426-RA 116 SYWLLEEEFMHIVFVHYLEVK 136
******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 79.701 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.7E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 9.8E-50 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.4E-7 | 391 | 508 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.3E-5 | 421 | 506 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 6.38E-16 | 421 | 506 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 1.0E-14 | 579 | 722 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 13.443 | 591 | 706 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.22E-14 | 592 | 699 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.19E-7 | 623 | 694 | No hit | No description |
| SuperFamily | SSF52540 | 2.42E-8 | 811 | 864 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.016 | 813 | 835 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.553 | 814 | 843 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.7E-4 | 815 | 834 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.006 | 836 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.523 | 837 | 861 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 3.6E-4 | 840 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 992 aa Download sequence Send to blast |
MANGGFINPG IRLDIQQLQL DARHRWLRAA EICEILRNHQ KFQISSEPPN QPSSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTIKEAHEK LKVDSVDMLH CYYAHGDDNE CFQRRSYWLL 120 EEEFMHIVFV HYLEVKGNRN SSRETEPVIS TSQISNSMSS SVSNSYNDAF TGNAGSPSAA 180 SSLNSYDEIE SEDNHQAISR YPSPSFSHVE ARFENNKMGA IINSTFLPSY QEEYKGPQPL 240 DYVSLVQQGV GTKDDNQFKT LDLASWEEVF QQSTRSSENL QANPLVSAAP LVGSTDVFNE 300 NFSLFLDAGF GVDSMEGGSL ASYLPNYNPL STEFPDNTNV SVIAKHPLLG NILMGEGLQK 360 VDSFSKWMSK EFSGVDDLYL NSSLDYSWQN IETSTSVDDP SIQHENYTVS PSIGYDQLFD 420 IQDFSPSCSS NDTETKVVIT GKFLVDPAEV FKYNWSCMFG EVEVPAEVLG NGILCCYAPL 480 HTAGRVPFYV TCSNRFACSQ VREFEYLLDS QRAINRSVSG DSIMEQQLWH RLQKLLSLKN 540 PANLDPVTKS FRKNEPIFSK ILLLMEDELC LGVEQQLKEK VVSWLFDKVS EDGKGPNVLD 600 ELGQGLLHLA AALGLDWIIP PTVATGVSVN FRDVCGWTAL HWAAFYGRET TVALLVALEA 660 ASGAVTDPSP EFPLGRTAAD LASANGFKGI SGFLAEHSLT AHLETLTMAD RKLDTPLDDS 720 MNKAVHTMRE NVATPGIEGD GSDLPLKDSL AAVRNATQAA GRIHQVFRMQ SFQRKHVSKG 780 AGPFGNYESL LSDEQLLLLV SSKIQRPGHP DEQTHSAAIR IQKKYRGYKK RKEFLIIRER 840 VVRIQAHVRG HQVRKRYKTV VWSVGILEKV ILRWRRRGTG LRGFRPDALY KASSTLGTQD 900 VQKISSEDDD YDFLKDGRKQ TEERLQKALV RVKSMVQYPD GRAQYRRILT AVEGFQKNQV 960 HENTRKNSED VPVEGEEDKI YVESLLDVTF E* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
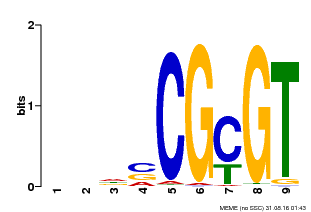
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010690491.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X1 | ||||
| Refseq | XP_010690492.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A0K9QYQ2 | 0.0 | A0A0K9QYQ2_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010690491.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.3 | 0.0 | ethylene induced calmodulin binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_017426-RA |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



