 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_015165-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 735aa MW: 80047.5 Da PI: 6.1727 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 49.4 | 7.8e-16 | 17 | 69 | 3 | 51 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNr 51
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNr
AHYPO_015165-RA 17 KYVRYTPEQVEALERLYHECPKPSSLRRQQLIRECpilsNIEPKQIKVWFQNR 69
5679************************************************9 PP
| |||||||
| 2 | START | 177.7 | 6.7e-56 | 115 | 323 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galql 93
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l
AHYPO_015165-RA 115 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTIEL 207
7899******************************************************.8888888888****************9999***** PP
EEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHH CS
START 94 mvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrs 183
+++l+a+++l+p Rdf+ +Ry+ +++g++v++++S++++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+
AHYPO_015165-RA 208 LYMQLYAPTTLAPaRDFWLLRYTSIMEDGSLVVCERSINNTQNGPSmpsVQNFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRP 301
*********************************************9888899****************************************** PP
HHHHHHHHHHHHHHHHTXXXXX CS
START 184 lvksglaegaktwvatlqrqce 205
l++s ++ ++kt++a+l+++++
AHYPO_015165-RA 302 LYESPTVVAQKTTMAALRQLRQ 323
******************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 12.892 | 12 | 69 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-9 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 7.27E-13 | 16 | 69 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.43E-13 | 17 | 69 | No hit | No description |
| Pfam | PF00046 | 2.0E-13 | 18 | 69 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-15 | 20 | 69 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50848 | 25.976 | 105 | 333 | IPR002913 | START domain |
| CDD | cd08875 | 2.68E-79 | 109 | 325 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 1.7E-22 | 114 | 320 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 2.33E-38 | 114 | 326 | No hit | No description |
| SMART | SM00234 | 5.3E-39 | 114 | 324 | IPR002913 | START domain |
| Pfam | PF01852 | 2.9E-53 | 115 | 323 | IPR002913 | START domain |
| Pfam | PF08670 | 1.8E-17 | 648 | 721 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 735 aa Download sequence Send to blast |
MMSMSCKDGK GGMDNGKYVR YTPEQVEALE RLYHECPKPS SLRRQQLIRE CPILSNIEPK 60 QIKVWFQNRS FFIWWLQSGI ASKDTSCESV VTSGQHHLTP QHPPRDASPA GLLSIAEETL 120 AEFLSKATGT AVEWVQMPGM KPGPDSIGIV AISHGCTGVA ARACGLVGLE PTRVAEILKD 180 RPSWFRDCRA VDVLNVLPTA NGGTIELLYM QLYAPTTLAP ARDFWLLRYT SIMEDGSLVV 240 CERSINNTQN GPSMPSVQNF VRAEMLPSGY LIRPCEGGGS IIHIVDHMDL EPWSVPEVLR 300 PLYESPTVVA QKTTMAALRQ LRQIAQEVSQ PNVSNWGRRP AALQALSQKL SRGFNEALNG 360 FTDEGWSMSG NDGVDDVTIL VNSSPDKLMA LNLAYANGFS SVSNTVLCAR ASMLLQNVPP 420 AVLLRFLREH RSEWADNNID AYLAAAAKIS PCSLPGSRMG GFGNQVILPL AHTIEHEELL 480 EVIKLEGVVS CPEDAMMPRD MFLLQLCSGM DENAVGTCSE LIFAPIDASF ADDAPLLPSG 540 FRIIPLDSVK ESSSPNRTLD LASALEVGPS ANRSSSDNSA SGNNMRSVMT IAFQFAFESH 600 LQENVASMAR QYVRSIISSV QRVALALSPH LGSQAGLRSP LGNPEAHTLA RWIVQSYRCF 660 LGVELLKSTG EGSDAILKSL WHHSDAIMCC SLKALPVFTF ANQAGFCMPS RRHLSFKHGP 720 SSFIREGSGV ESAE* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
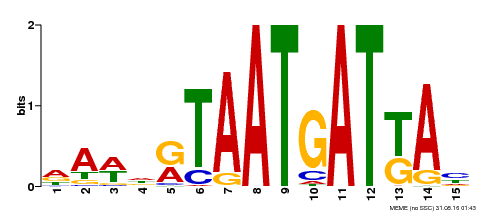
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010691423.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A0K9RV39 | 0.0 | A0A0K9RV39_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010691422.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.3 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_015165-RA |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



