 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_007032-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 555aa MW: 62781.7 Da PI: 7.2479 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.3 | 4e-26 | 88 | 189 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtklel 83
fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k e++ +r+ +t+Cka+++vk++ dg+w+++++ +
AHYPO_007032-RA 88 FYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKGnrprirqcnkqePESATGRRSCGKTNCKASMHVKRRPDGRWVIHSFIK 181
9**************************************999998889999999775555555589999************************* PP
FAR1 84 eHnHelap 91
eHnHel p
AHYPO_007032-RA 182 EHNHELLP 189
*****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.0E-24 | 88 | 189 | IPR004330 | FAR1 DNA binding domain |
| PROSITE profile | PS50966 | 8.864 | 274 | 310 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 5.9E-5 | 284 | 309 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 4.4E-8 | 285 | 312 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 555 aa Download sequence Send to blast |
MDIDLRLPSG VQDRQDEDEN AVSDMLDRED KLDDGVDAED ALHAGEVHVE DELTLNSTIS 60 DMGDFKEDAN LEPLAGMEFV SHQEAYAFYQ EYARSMGFNT AIQNSRRSKT SREFIDAKFA 120 CSRYGTKREY DKGNRPRIRQ CNKQEPESAT GRRSCGKTNC KASMHVKRRP DGRWVIHSFI 180 KEHNHELLPA QAVSEQTRKM YAAMARNYAE YKNVVALKSP SPLEKHLSTL YTHAVFKKFQ 240 VEVLGAVACH PKTERQDEVY TTFRVQDFEK NQEFFITWNP MKSEVSCMCH LFEYKGYLCR 300 HAMVVFQMCG LSMIPSQYIL KRWKKDAKVR HLIVEGTEQV PSRVQRYNDL CNRAMRLGEE 360 GSLSQESYTI ALRALDEAFN NCIGVNNVCK TLPEVSTSAN HGLLCIEDDS PSRSFSKTIK 420 KKHPTKKRKI NTEAEAMTIG SQYNLQQMDK LGSRPVRPLD GFYGPQQGVP GMVQLNLMAP 480 TDNYYGNQPA IQGLGQLNSI APSNDGYYGQ PGMHGLGQSQ MDIFRSPTGF SYGIRDEANV 540 RSAQLHDDGG PRHS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
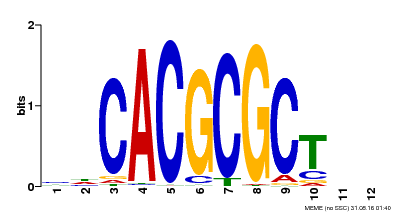
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010680157.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 1e-125 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A0A0KV81 | 0.0 | A0A0A0KV81_CUCSA; Uncharacterized protein | ||||
| STRING | XP_010680156.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 1e-125 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_007032-RA |



