 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AHYPO_005576-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Amaranthaceae; Amaranthus
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 408aa MW: 44847.5 Da PI: 9.5076 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.1 | 1e-13 | 79 | 127 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+++++g
AHYPO_005576-RA 79 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDVAAQRFRG 127
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 94.6 | 7e-30 | 211 | 322 | 1 | 93 |
EEEE-..-HHHHTT-EE--HHH.HTT...................---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT-- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh...................ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLk 75
f+kv tpsdv+k++rlv+pk++ae+h ++++ +++ l +ed + ++W+++++y+++s++yvltkGW++Fvk++ Lk
AHYPO_005576-RA 211 FEKVVTPSDVGKLNRLVIPKQHAEKHfplqannnnngngilfstsSNNSSKGVLLNFEDVNEKVWRFRYSYWNSSQSYVLTKGWSRFVKEKSLK 304
99****************************9999999999888886666799****************************************** PP
TT-EEEEEE-SSSEE..E CS
B3 76 egDfvvFkldgrsefelv 93
+gD+v F+++ +++l+
AHYPO_005576-RA 305 AGDVVSFQRSTGPDKQLY 322
********7755666555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 3.2E-9 | 79 | 127 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.77E-17 | 79 | 135 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 4.58E-27 | 79 | 134 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 8.6E-21 | 80 | 135 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.455 | 80 | 135 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.6E-30 | 80 | 141 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 1.1E-38 | 207 | 328 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 8.89E-29 | 209 | 319 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.57E-26 | 210 | 314 | No hit | No description |
| Pfam | PF02362 | 3.2E-27 | 211 | 322 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.0E-24 | 211 | 327 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.169 | 211 | 329 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 408 aa Download sequence Send to blast |
MDSSSPSSMD ECSSTSDSLT STTPPLPPTL IRTLIPTAPF KASAAAAKPG SLCRMGSGTS 60 VIIDSENGVE AESRKLPSSK YKGVVPQPNG RWGAQIYEKH QRVWLGTFNE EDEAARAYDV 120 AAQRFRGRDA VTNFKPLSAE DEAGSAELRF LNSRSKAEVV DMLRKHTYLD ELQQSKRNYK 180 ADFDKTGSTH LISDKSKPEN TINGFGKEQL FEKVVTPSDV GKLNRLVIPK QHAEKHFPLQ 240 ANNNNNGNGI LFSTSSNNSS KGVLLNFEDV NEKVWRFRYS YWNSSQSYVL TKGWSRFVKE 300 KSLKAGDVVS FQRSTGPDKQ LYIDYKQRNC DGLLGRVMEG PVNVGSTGQM LRLFGVDICK 360 VHHPIVVNVV ADGNGNGNGC GNGKRLREVE LLAMESNSKK QMVVAAL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 5e-55 | 208 | 333 | 11 | 122 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
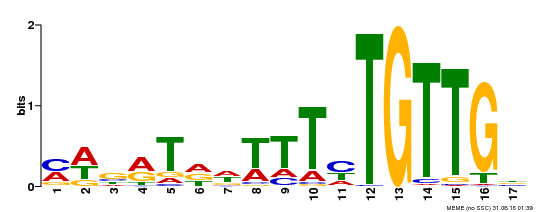
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021845494.1 | 0.0 | AP2/ERF and B3 domain-containing transcription factor RAV1-like | ||||
| Swissprot | Q9ZWM9 | 1e-138 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | A0A0K9QUD7 | 0.0 | A0A0K9QUD7_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010681667.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-137 | related to ABI3/VP1 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AHYPO_005576-RA |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



