 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.2816s0008.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 253aa MW: 28783.7 Da PI: 8.6585 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 99.7 | 1.1e-31 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krienk+nrqvtf+kRrng+lKKA+ELSvLCdaeva+iifs++gklye++s
Araha.2816s0008.1.p 9 KRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLYEFCS 59
79***********************************************96 PP
| |||||||
| 2 | K-box | 97.5 | 2.1e-32 | 78 | 173 | 4 | 99 |
K-box 4 ssgksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenka 93
+ +++ +++ e++++e+ kLk ++enLqr+qR+llGedL++L+ keL+qLe+qL+ slk++Rs K++++l+q+++lq+ke++l e+n+a
Araha.2816s0008.1.p 78 IEVNNKPAKELENSYREYLKLKGRYENLQRQQRNLLGEDLGPLNSKELEQLERQLDGSLKQVRSIKTQYMLDQLSDLQNKEQMLLETNRA 167
56667788999******************************************************************************* PP
K-box 94 Lrkkle 99
L +kl+
Araha.2816s0008.1.p 168 LAMKLD 173
**9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 1.3E-41 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 33.515 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 3.66E-34 | 2 | 80 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 3.48E-45 | 2 | 77 | No hit | No description |
| PRINTS | PR00404 | 9.4E-33 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.4E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.4E-33 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.4E-33 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 9.4E-26 | 85 | 172 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 14.889 | 88 | 178 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 253 aa Download sequence Send to blast |
MGRGRVELKR IENKINRQVT FAKRRNGLLK KAYELSVLCD AEVALIIFSN RGKLYEFCSS 60 SNMLKTLDRY QKCSYGSIEV NNKPAKELEN SYREYLKLKG RYENLQRQQR NLLGEDLGPL 120 NSKELEQLER QLDGSLKQVR SIKTQYMLDQ LSDLQNKEQM LLETNRALAM KLDDMIGVRS 180 HHMGGGGGVW EGGEQNVTYA HHQVQSQGLY QPLECNPTLQ MGYDNPVCSE QITATTQAQA 240 QQGNGYIPGW ML* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ox0_A | 2e-34 | 75 | 174 | 1 | 103 | Developmental protein SEPALLATA 3 |
| 4ox0_B | 2e-34 | 75 | 174 | 1 | 103 | Developmental protein SEPALLATA 3 |
| 4ox0_C | 2e-34 | 75 | 174 | 1 | 103 | Developmental protein SEPALLATA 3 |
| 4ox0_D | 2e-34 | 75 | 174 | 1 | 103 | Developmental protein SEPALLATA 3 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Functions with SEPALLATA2/AGL4 and SEPALLATA3/AGL9 to ensure proper development of petals, stamens and carpels, and to prevent the indeterminate growth of the flower meristem. Forms a heterodimer via the K-box domain with AGAMOUS, that could be involved in genes regulation during floral meristem development. | |||||
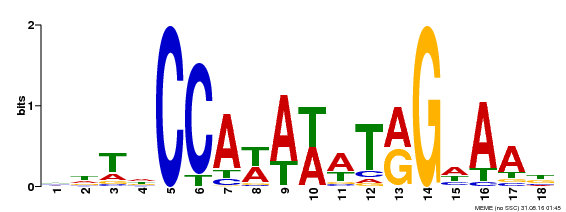
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00092 | SELEX | Transfer from AT5G15800 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.2816s0008.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK118608 | 0.0 | AK118608.1 Arabidopsis thaliana At5g15800 mRNA for putative transcription factor (AGL2), complete cds, clone: RAFL19-86-A21. | |||
| GenBank | BT006224 | 0.0 | BT006224.1 Arabidopsis thaliana At5g15800 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006400104.1 | 1e-176 | developmental protein SEPALLATA 1 | ||||
| Swissprot | P29382 | 1e-166 | SEP1_ARATH; Developmental protein SEPALLATA 1 | ||||
| TrEMBL | A0A178UQB3 | 1e-179 | A0A178UQB3_ARATH; SEP1 | ||||
| TrEMBL | Q5XXN7 | 1e-179 | Q5XXN7_ARATH; SEPALLATA1 | ||||
| STRING | XP_006400104.1 | 1e-176 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3289 | 27 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G15800.1 | 1e-168 | MIKC_MADS family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.2816s0008.1.p |



