 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.19926s0004.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 736aa MW: 80805.1 Da PI: 5.9853 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.7 | 6.1e-21 | 69 | 124 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Araha.19926s0004.1.p 69 KKRYHRHTPQQIQELESMFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 124
688999***********************************************999 PP
| |||||||
| 2 | START | 213.1 | 1e-66 | 257 | 476 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....ka 79
ela++a++elvk+a +eep+Wvks +++n+de++++f+++k +ea+r+sg+v+ ++ lve+l+d++ +W+e+++ +a
Araha.19926s0004.1.p 257 ELALTAMDELVKLAHSEEPLWVKSLdgerDELNQDEYMRTFSSTKPtgLATEASRTSGMVIINSLALVETLMDSN-RWTEMFPcnvaRA 344
5899*************************************9999999***************************.************* PP
EEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCE CS
START 80 etlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksngh 161
+t++vissg galqlm+aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+++ ++ + ++v+R +lpSg++++++sng+
Araha.19926s0004.1.p 345 TTTDVISSGmagtinGALQLMNAELQVLSPLVPvRNVNFLRFCKQHAEGVWAVVDVSIEPVRESSGGAPVIR--RLPSGCVVQDMSNGY 431
****************************************************************99******..*************** PP
EEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 162 skvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
skvtwveh++++++++h+l+r+l++sgl +g ++w+atlqrqc++
Araha.19926s0004.1.p 432 SKVTWVEHAEYDENHIHQLYRPLLRSGLGFGSQRWLATLQRQCQC 476
*******************************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.5E-20 | 56 | 126 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-22 | 57 | 126 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.297 | 66 | 126 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.5E-18 | 67 | 130 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.83E-18 | 68 | 126 | No hit | No description |
| Pfam | PF00046 | 1.8E-18 | 69 | 124 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 101 | 124 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 42.27 | 248 | 479 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.84E-29 | 252 | 476 | No hit | No description |
| CDD | cd08875 | 1.79E-112 | 252 | 475 | No hit | No description |
| Pfam | PF01852 | 4.3E-60 | 257 | 476 | IPR002913 | START domain |
| SMART | SM00234 | 1.4E-40 | 257 | 476 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.83E-21 | 495 | 720 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 736 aa Download sequence Send to blast |
MFGQEQPERG TNRGEASMRN NNVGGGDNFD GSSVNRRSRE EEHESRSGSD NVEGISGEDQ 60 DAADKPPRKK RYHRHTPQQI QELESMFKEC PHPDEKQRLE LSKRLCLETR QVKFWFQNRR 120 TQMKTQLERH ENALLRQEND KLRAENMSIR EAMRNPICTN CGGPAMLGDV SLEEHHLRIE 180 NARLKDELDR VCNLTGKFLG HHHHHYNSSL ELAVGTNNGG NFAFPPDFGG GGGCLPPPQQ 240 QQPTGIDGID QKSVLLELAL TAMDELVKLA HSEEPLWVKS LDGERDELNQ DEYMRTFSST 300 KPTGLATEAS RTSGMVIINS LALVETLMDS NRWTEMFPCN VARATTTDVI SSGMAGTING 360 ALQLMNAELQ VLSPLVPVRN VNFLRFCKQH AEGVWAVVDV SIEPVRESSG GAPVIRRLPS 420 GCVVQDMSNG YSKVTWVEHA EYDENHIHQL YRPLLRSGLG FGSQRWLATL QRQCQCLAIL 480 MSSSVTSHDN TSITPGGRKS MLKLAQRMTF NFCSGISAPS VHSWSKLTVG NVDPDVRVMT 540 RKSVDDPGEP PGIVLSAATS VWLPASPQRL YDFLRNERMR CEWDILSNGG PMQEMAHIAK 600 GQDQGVSLLR SNAMNANQSS MLILQETCID ASGALVVYAP VDIPAMHVVM NGGDSSYVAL 660 LPSGFAVLPD GGIDGGGSGD GEQRPDGGGS LLTVAFQILV NNLPTAKLTV ESVETVNNLI 720 SCTVQKIRAA LQCES* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
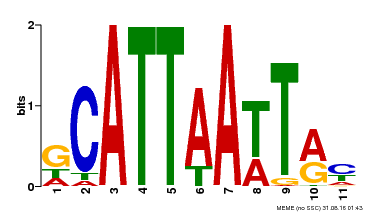
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.19926s0004.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK226968 | 0.0 | AK226968.1 Arabidopsis thaliana mRNA for homeodomain protein AHDP, complete cds, clone: RAFL09-36-B10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020878511.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | R0H8W0 | 0.0 | R0H8W0_9BRAS; Uncharacterized protein | ||||
| STRING | Bostr.10064s0036.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.19926s0004.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



