 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.1729s0004.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 485aa MW: 52553.5 Da PI: 4.8458 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 82.2 | 4.4e-26 | 167 | 232 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf++l++++++g+++ln++a++L +v ++RRiYDi+NVLe+++liek kn+i wkg
Araha.1729s0004.1.p 167 RYDSSLGLLTKKFVNLIKQAKDGMLDLNKAAETL---EV--QKRRIYDITNVLEGIDLIEKPFKNRILWKG 232
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 8.0E-28 | 163 | 234 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 1.63E-16 | 167 | 230 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 2.2E-32 | 167 | 232 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| Pfam | PF02319 | 9.1E-23 | 169 | 232 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 3.30E-49 | 244 | 348 | No hit | No description |
| SuperFamily | SSF144074 | 2.38E-32 | 244 | 348 | No hit | No description |
| Pfam | PF16421 | 1.3E-32 | 247 | 346 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051446 | Biological Process | positive regulation of meiotic cell cycle | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 485 aa Download sequence Send to blast |
MSGLVRSSPG SSHPPPPPPH HPPSSPAPVT STPVVPPIRR HLAFASTKPP FHPSDDYHRF 60 NPSSLTNNND RSFVNACGVV DREEDAVVVR SPSRKRKSTM DMVVAPSNNG FTSSGFTSIP 120 SSPCQTPAKG GRVNIKSKAK GNKSTPQTPI STNAGSPVTL TPSGSCRYDS SLGLLTKKFV 180 NLIKQAKDGM LDLNKAAETL EVQKRRIYDI TNVLEGIDLI EKPFKNRILW KGVDASPGDE 240 DADVSVLQAE IENLALEEQA LDNQIRQTEE RLRDLSENET NQKWLFVTEE DIKSLPGFQN 300 QTLIAVKAPH GTTLEVPDPD EAVDHPQRRY RIILRSTMGP IDVYLVSEFE GKFEDTNGSV 360 AAAPACLPIA SSSGSTGHHD IEALTVDNTG TAIEHQVSHD HPQPGDTSEL NYLQEQVGGM 420 LKITPSDVEN DESDYWLLSS AEISMTDIWK TDSGIDWDYG IADVSTPPPG MGEIAPTAVD 480 STPR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 3e-23 | 162 | 233 | 2 | 74 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 92 | 97 | SRKRKS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Regulates gene expression both positively and negatively. Activates the expression of E2FB. Involved in the control of cell-cycle progression from G1 to S phase. Stimulates cell proliferation and delays differentiation. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11889041, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:15377755, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:19662336}. | |||||
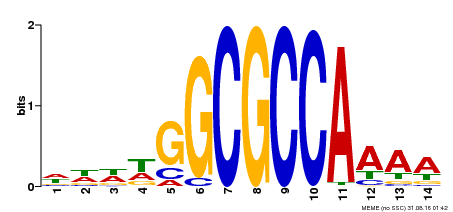
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00293 | DAP | Transfer from AT2G36010 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.1729s0004.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF242582 | 0.0 | AF242582.1 Arabidopsis thaliana E2F transcription factor-3 E2F3 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002879589.1 | 0.0 | transcription factor E2FA | ||||
| Swissprot | Q9FNY0 | 0.0 | E2FA_ARATH; Transcription factor E2FA | ||||
| TrEMBL | D7LIM9 | 0.0 | D7LIM9_ARALL; E2F transcription factor-3 | ||||
| STRING | fgenesh2_kg.4__1634__AT2G36010.3 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM808 | 28 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G36010.1 | 0.0 | E2F transcription factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.1729s0004.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



