 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.16804s0008.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 271aa MW: 30104.2 Da PI: 9.8991 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 304 | 5.5e-93 | 9 | 270 | 15 | 301 |
GAGA_bind 15 epaaslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllvenslasalpvgvqvls 103
e++++l +++++ms++aerda+++e n a++++k+a+a+rd+a+ qrdkal+er++a++e +++l al ++en+l+ l+ +
Araha.16804s0008.1.p 9 EQHNALV--MNKKIMSILAERDAAVKEINEAVAATKEALAARDEALEQRDKALSERDNAIMETESALNALRYRENNLNYILSC--A-KR 92
4556677..99****************************************************************96654422..1.22 PP
GAGA_bind 104 gtksidslqqlsepqledsavelreeeklealpieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkkvkkesaderskaekk 192
g ++ + +++ pi++ ++e +++ +k r+k+ k+ ++k kk +e+ +++v++++ ++s+++++
Araha.16804s0008.1.p 93 GGSHSC---------------VTDKSHLPAPSPISTIPPEPANTRLTK---RKKESKQVARSKGKKVGEDLNHQVASPG--KKSRKDWD 161
222222...............123445556788999999888888777...4455555566667777889999*****9..57****** PP
GAGA_bind 193 sidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstkrrgaRiagrKmSqgafkklLekLaaeGydlsn 281
s d++ln v++De+t+PvP+C+CtG+ rqCYkWGnGGWqS+CCttt+S+yPLP+++++r++R+agrKmS+++f++lL++La+eG++ls
Araha.16804s0008.1.p 162 SYDVGLNLVTFDETTMPVPMCTCTGTARQCYKWGNGGWQSSCCTTTLSQYPLPQMPNKRHSRVAGRKMSGSVFSRLLSRLAGEGHELSS 250
***************************************************************************************** PP
GAGA_bind 282 pvDLkdhWAkHGtnkfvtir 301
vDLkd+WA+HGtn+++ti+
Araha.16804s0008.1.p 251 HVDLKDYWARHGTNRYITIK 270
*******************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 2.0E-140 | 1 | 270 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.6E-89 | 8 | 270 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 271 aa Download sequence Send to blast |
MVPQHQIKEQ HNALVMNKKI MSILAERDAA VKEINEAVAA TKEALAARDE ALEQRDKALS 60 ERDNAIMETE SALNALRYRE NNLNYILSCA KRGGSHSCVT DKSHLPAPSP ISTIPPEPAN 120 TRLTKRKKES KQVARSKGKK VGEDLNHQVA SPGKKSRKDW DSYDVGLNLV TFDETTMPVP 180 MCTCTGTARQ CYKWGNGGWQ SSCCTTTLSQ YPLPQMPNKR HSRVAGRKMS GSVFSRLLSR 240 LAGEGHELSS HVDLKDYWAR HGTNRYITIK * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
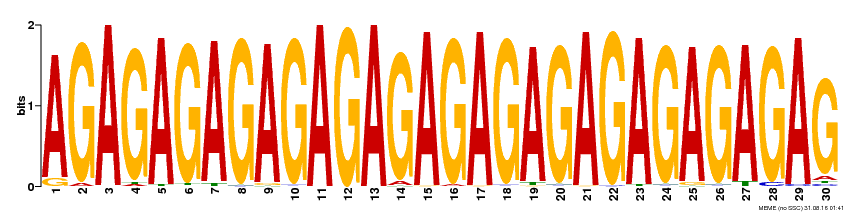
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00480 | DAP | Transfer from AT4G38910 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.16804s0008.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF633603 | 0.0 | EF633603.1 Cardamine pratensis GAGA-motif binding transcriptional activator (BBR/BPC5) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020875421.1 | 0.0 | protein BASIC PENTACYSTEINE5 isoform X1 | ||||
| Refseq | XP_020875422.1 | 0.0 | protein BASIC PENTACYSTEINE5 isoform X2 | ||||
| Swissprot | F4JUI3 | 1e-171 | BPC5_ARATH; Protein BASIC PENTACYSTEINE5 | ||||
| TrEMBL | V4M9X7 | 0.0 | V4M9X7_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006411681.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4090 | 26 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G38910.1 | 1e-165 | basic pentacysteine 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.16804s0008.1.p |



