 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aradu.HL1PG | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 843aa MW: 93334.7 Da PI: 6.6696 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.8 | 5e-24 | 151 | 252 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f+k+lt sd++++g +++ +++a+e+ + k++ +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +++el
Aradu.HL1PG 151 FCKTLTASDTSTHGGFSVLRRHADEClppldmS-KQPPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGENGEL 245
99*****************************63.4445569************************************************..449999* PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+v+r+
Aradu.HL1PG 246 RVGVRRA 252
****996 PP
| |||||||
| 2 | Auxin_resp | 113.2 | 2.5e-37 | 277 | 359 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+ t+++F+v+Y+Pr+s++eF+v++++++++lk+++++G Rfkm+fe+e+++e+r++Gt+vg++d+d+ rWp+SkWrsLk
Aradu.HL1PG 277 AWHAILTGTMFTVYYKPRTSPAEFIVPYDQYMESLKNNYTIGLRFKMRFEGEEAPEQRFTGTIVGIEDADSKRWPESKWRSLK 359
79*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.0E-41 | 147 | 266 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.26E-40 | 147 | 280 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.61E-22 | 149 | 251 | No hit | No description |
| Pfam | PF02362 | 5.6E-22 | 151 | 252 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.8E-23 | 151 | 253 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.026 | 151 | 253 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.0E-35 | 277 | 359 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 22.309 | 717 | 799 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 3.34E-9 | 719 | 794 | No hit | No description |
| Pfam | PF02309 | 9.8E-8 | 759 | 804 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MASTEELMNR KVETNNNNGG MNAAEAQNAS SSSSSSSARD AEAALYKELW HACAGPLVTV 60 PKEGELVFYF PQGHIEQVEA STNQVAEQHM PVYDLPSKIL CRVINVMLKA EPDTDEVFAQ 120 LTLLPEPNQD ENAVEKAIPP APPPRFHVHS FCKTLTASDT STHGGFSVLR RHADECLPPL 180 DMSKQPPTQE LVAKDLHGNE WRFRHIFRGQ PRRHLLQSGW SVFVSSKRLV AGDAFIFLRG 240 ENGELRVGVR RAMRQQGNVP SSVISSHSMH LGVLATAWHA ILTGTMFTVY YKPRTSPAEF 300 IVPYDQYMES LKNNYTIGLR FKMRFEGEEA PEQRFTGTIV GIEDADSKRW PESKWRSLKV 360 RWDETSTIPR PERVSPWKIE PALAPPALNP LPMPRPKRPR SNAIPSSPDS SVLTREASSK 420 ASVDPLPPSG FPRVLQGQEF STLRGNFAES NESDTAEKSV VWPPAVDDEK IDVVSTSRRY 480 GPESWMSMGR HELPYSDLLS GFGANGDPSS RSSLIDQSDS VANPGRKHLL DREGKTNVPS 540 SSPWSVIPSS LSLNLLEPNA KVSAQGGDAA YQIRGNLRYG AFGEYPVLHG HKVENSHGNL 600 SMPPPPLAQY ESPRSKELLR KPKSAKTSDV ARPKDGDCKL FGISLLSSPP APETPTSQRN 660 VASETVSHTY PTHQHRTFEN EQKSENSMGS KATDGLAVVD ELHLRDVQPK PHSGSARSCT 720 KVHKKGIALG RSVDLTKFSG YGELIAELDQ LFEFGGELIS AKKDWLTVYT DNEGDMMLVG 780 DDPWQEFCAM VRKIYIYPRE EIQKMSPGTL SSKNEENQLA SEGVDAQEIR CQQHHPTSTP 840 DHS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-178 | 29 | 381 | 3 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-178 | 29 | 381 | 3 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-178 | 29 | 381 | 3 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-178 | 29 | 381 | 3 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-178 | 29 | 381 | 3 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
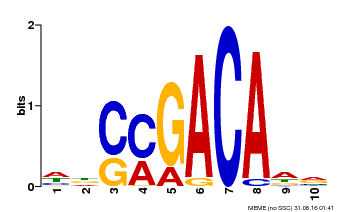
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Aradu.HL1PG |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015955707.1 | 0.0 | auxin response factor 2 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A445DTD0 | 0.0 | A0A445DTD0_ARAHY; Auxin response factor | ||||
| STRING | GLYMA05G38540.4 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



