 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aradu.9MQ2E | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 320aa MW: 36410.9 Da PI: 6.527 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.3 | 7.5e-08 | 43 | 88 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqk 46
+WT++E +++ +a +++ ++ +W ++a++++ g+t +++ ++
Aradu.9MQ2E 43 KWTPQENKQFENALALFDKEtpdRWLKVAAMIP-GKTVGDVIKQYRE 88
8*****************99*************.*******999986 PP
| |||||||
| 2 | Myb_DNA-binding | 47.5 | 4.2e-15 | 151 | 195 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + k++G+g+W+ I+r + ++Rt+ q+ s+ qky
Aradu.9MQ2E 151 PWTEEEHRLFLMGLKKYGKGDWRNISRNFVTTRTPTQVASHAQKY 195
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 10.182 | 39 | 94 | IPR017884 | SANT domain |
| SMART | SM00717 | 4.8E-9 | 40 | 92 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.65E-11 | 42 | 98 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.1E-6 | 43 | 89 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.68E-7 | 43 | 90 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-13 | 142 | 201 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.858 | 144 | 200 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.66E-17 | 145 | 199 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-17 | 147 | 199 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.7E-13 | 148 | 198 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.7E-12 | 151 | 195 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.67E-11 | 151 | 196 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 320 aa Download sequence Send to blast |
MYATIGYSLL EEEEEMNRGI EVLSPASYLH NSNWLFQESA GTKWTPQENK QFENALALFD 60 KETPDRWLKV AAMIPGKTVG DVIKQYRELV EDVNVIESGF IPIPGGTTDG SFTLEWVDNH 120 EFDGFKQFYG VGCKRGSSTR PSEQERKKGV PWTEEEHRLF LMGLKKYGKG DWRNISRNFV 180 TTRTPTQVAS HAQKYFIRQL NGGKDKRRSS IHDITMVNLP ETKSPSSESN KPPSPDANLN 240 QSQKLLSSMV KQEYDWKSPP YEGVPLVFDS TNGNSFMTPL CGASSYGSKP QEQYLLRGTF 300 HGFQFGPYDT FMQMQPMQPQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 4e-16 | 40 | 110 | 7 | 77 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
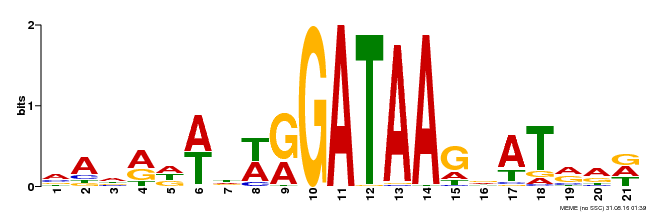
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Aradu.9MQ2E |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015950297.1 | 0.0 | transcription factor DIVARICATA | ||||
| Swissprot | Q8S9H7 | 1e-116 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A445AK09 | 0.0 | A0A445AK09_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA03G14440.1 | 1e-171 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF466 | 34 | 162 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 1e-112 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



