 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aqcoe7G131800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Ranunculales; Ranunculaceae; Thalictroideae; Aquilegia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 312aa MW: 34219.6 Da PI: 9.0967 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.9 | 5.4e-14 | 106 | 150 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ ++ + ++lG+g+W+ I++ + ++Rt+ q+ s+ qky
Aqcoe7G131800.1.p 106 PWTEEEHRTFLAGLAKLGKGDWRGISKNFVTTRTPTQVASHAQKY 150
7*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 19.02 | 99 | 155 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.24E-17 | 101 | 154 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.1E-17 | 102 | 154 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.5E-10 | 103 | 153 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.3E-12 | 104 | 150 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.7E-12 | 106 | 150 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.01E-9 | 106 | 151 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 312 aa Download sequence Send to blast |
MGRKCSHCGN MGHNLRTCTT YRGGGGGGGV GGVRLFGVQL DIASSLPMKK SFSMSCLSPT 60 SSTSSSSPSP LSSSCRPVPE SLDKTSNGYL SDGLISRPLE KKKGIPWTEE EHRTFLAGLA 120 KLGKGDWRGI SKNFVTTRTP TQVASHAQKY FLRQISLNTK KRRSSLFDMV ESNKMDKHHV 180 NRSCSFKTHD SSSMSFKLHS VPTQPFDDNK TTVQGKVHED ITLHLIDLNL VEQDCRLDNQ 240 GTEKLQFRQF LHLMPVWAYE SVDDYSTSST PTKLISSNAA SDGQELTLEP SMSLDRNKSC 300 PDSFTSGTIL V* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds to 5'-TATCCA-3' elements in gene promoters. Contributes to the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Transcription repressor involved in a cold stress response pathway that confers cold tolerance. Suppresses the DREB1-dependent signaling pathway under prolonged cold stress. DREB1 responds quickly and transiently while MYBS3 responds slowly to cold stress. They may act sequentially and complementarily for adaptation to short- and long-term cold stress (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
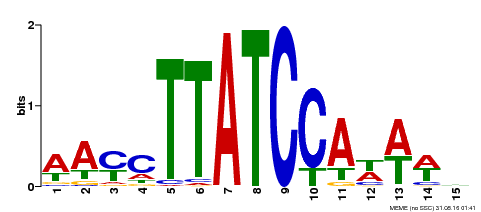
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA) (PubMed:12172034). Induced by cold stress in roots and shoots. Induced by salt stress in shoots. Down-regulated by abscisic aci (ABA) in shoots (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010262182.1 | 1e-89 | PREDICTED: transcription factor MYB1R1-like | ||||
| Swissprot | Q7XC57 | 6e-50 | MYBS3_ORYSJ; Transcription factor MYBS3 | ||||
| TrEMBL | A0A2G5F8Q8 | 0.0 | A0A2G5F8Q8_AQUCA; Uncharacterized protein | ||||
| STRING | Aquca_001_00103.1 | 0.0 | (Aquilegia coerulea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56840.1 | 8e-54 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aqcoe7G131800.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



