 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco016874.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 317aa MW: 34845 Da PI: 9.1716 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 37.3 | 6.4e-12 | 31 | 77 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqky 47
rWT eE +l+ da +++ g+ W+++a+ ++ g+t +++s+++++
Aco016874.1 31 RWTREENKLFEDALARFEGDsqdKWEKVAAAIP-GKTVGDVVSHYRNL 77
9********************************.************97 PP
| |||||||
| 2 | Myb_DNA-binding | 46.3 | 9.7e-15 | 142 | 186 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + +++G+g+W++I+r + +Rt+ q+ s+ qky
Aco016874.1 142 PWTEEEHRLFLMGLQKYGKGDWRSISRNFVMTRTPTQVASHAQKY 186
8*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 10.954 | 27 | 82 | IPR017884 | SANT domain |
| SMART | SM00717 | 2.5E-8 | 28 | 80 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.6E-5 | 30 | 78 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 3.8E-9 | 31 | 77 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.32E-8 | 31 | 78 | No hit | No description |
| SuperFamily | SSF46689 | 1.57E-10 | 31 | 85 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.768 | 135 | 191 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.5E-17 | 137 | 191 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.4E-18 | 138 | 189 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 5.6E-13 | 139 | 185 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.8E-13 | 139 | 189 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.1E-13 | 142 | 186 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.43E-11 | 142 | 187 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MESWMEVLPP AATDPFLCST VGGFGPRRSC RWTREENKLF EDALARFEGD SQDKWEKVAA 60 AIPGKTVGDV VSHYRNLVAD VRQIEAGLVP FPGYNSSAPP FTLEWEKQPY SSCLLGVGGR 120 RSSSSSSSGR SSSDQERKKG VPWTEEEHRL FLMGLQKYGK GDWRSISRNF VMTRTPTQVA 180 SHAQKYFIRL NSGGKDKRRS SIHDITTVNL PDSRPQSPSQ PSILGAKSNL AAPPPMPGHI 240 SPTIDANKQH HEAANNAFNS CAHTTNKYSM QPSFGASPYG TKLHPQISQC GPLNDSMVGD 300 HSMLFQMQFS QPHSHG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 8e-14 | 32 | 98 | 11 | 77 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
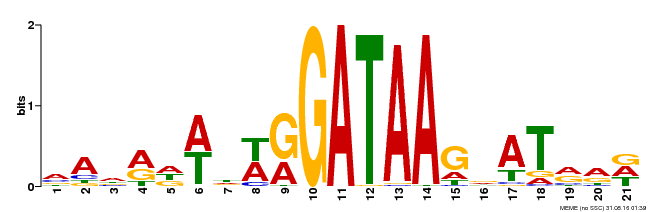
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM665060 | 2e-44 | KM665060.1 Ananas bracteatus clone 50154 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020086210.1 | 0.0 | transcription factor DIVARICATA-like | ||||
| Refseq | XP_020086211.1 | 0.0 | transcription factor DIVARICATA-like | ||||
| Refseq | XP_020086212.1 | 0.0 | transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 1e-88 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A199US12 | 0.0 | A0A199US12_ANACO; Transcription factor DIVARICATA | ||||
| STRING | XP_008785419.1 | 1e-113 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7886 | 33 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 1e-76 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco016874.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



