 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco013950.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 691aa MW: 74529.4 Da PI: 7.5421 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.9 | 4e-23 | 127 | 228 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefe 91
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g +W++++iyr++++r++lt+GW+ Fv++++L +gD++vF +++++
Aco013950.1 127 FAKTLTQSDANNGGGFSVPRYCAETIfprldysTDPPVQ--TVLAKDVHGVVWKFRHIYRGTPRRHLLTTGWSTFVNQKKLVAGDSIVFL--RTENGD 220
89****************************996444444..8999*********************************************..458899 PP
.EEEEE-S CS
B3 92 lvvkvfrk 99
l+v+++r+
Aco013950.1 221 LCVGIRRA 228
9****997 PP
| |||||||
| 2 | Auxin_resp | 119.2 | 3.2e-39 | 299 | 382 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a+++++FevvY+Prast+eF+vk+ v++a++v++++GmRfkmafetedss++++ +Gt+++v+ +dp+rWpnS+Wr+L+
Aco013950.1 299 AANLAASGQPFEVVYYPRASTPEFCVKAGAVRAAMRVQWCPGMRFKMAFETEDSSRISWfMGTISSVQVADPIRWPNSPWRLLQ 382
7899******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 3.3E-37 | 120 | 231 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.75E-29 | 122 | 233 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.77E-21 | 126 | 227 | No hit | No description |
| SMART | SM01019 | 6.4E-21 | 127 | 229 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.7E-21 | 127 | 228 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.83 | 127 | 229 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.0E-34 | 299 | 382 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 20.153 | 604 | 682 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 691 aa Download sequence Send to blast |
MITFTDSPKE AMKETEKCLD PQLWHACAGG MVQMPPIHSK VYYFPQGHAE HAQGGPADPL 60 GPSPSRVPPL ILARVSAIKF MADPETDEVF ARIRLVPVRP NEPDYGDHCE IGGNGDGPDA 120 QEKPASFAKT LTQSDANNGG GFSVPRYCAE TIFPRLDYST DPPVQTVLAK DVHGVVWKFR 180 HIYRGTPRRH LLTTGWSTFV NQKKLVAGDS IVFLRTENGD LCVGIRRAKR GVGAGPESPP 240 GWNTPNGNCV SAYGNFSVFL REEESKLMRG PANGGNMNGG GGGSLRGKGR VRAESVVEAA 300 NLAASGQPFE VVYYPRASTP EFCVKAGAVR AAMRVQWCPG MRFKMAFETE DSSRISWFMG 360 TISSVQVADP IRWPNSPWRL LQVTWDEPDL LQNVKRVSPW LVELVSNMPT IHLPPFSPPR 420 KKLCVPQHPG GFPLEAQLQG PAFHGSPFGP VSSPLCCFPD SAPAGIQGAR HAHLGLSISD 480 LHLSKLQSSL FAAAPPARIS TGLIVSNSPI RENISCLLTI GSPSQPTKKS VNGKTPQLVL 540 FGQPILTEQQ ISLSNSADTA SPGITKNSSS DGNPEKTVNV SDGSGSAVNQ NGHAENSSCE 600 ELVAGHCKVF MDSEDVGRTL DLSVLGSYEE LYRRLGDMFG IDRAELMSHV RYRDAAGAVK 660 HTGDEPFSEF MRSARRLTIV TDAGSDNLGR * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldy_A | 2e-69 | 22 | 403 | 22 | 353 | Auxin response factor 1 |
| 4ldy_B | 2e-69 | 22 | 403 | 22 | 353 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
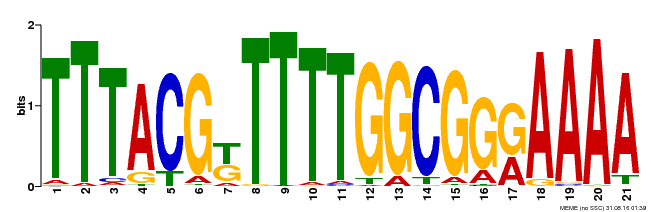
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020084645.1 | 0.0 | auxin response factor 18-like isoform X1 | ||||
| Refseq | XP_020084646.1 | 0.0 | auxin response factor 18-like isoform X1 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A199W052 | 0.0 | A0A199W052_ANACO; Auxin response factor | ||||
| STRING | XP_008791554.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP516 | 37 | 152 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco013950.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



