 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco006706.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 377aa MW: 39788.6 Da PI: 10.2664 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 40.2 | 8.7e-13 | 64 | 112 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f + eAa+a++ a+++++g
Aco006706.1 64 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNAEAEAARAYDVAAQRFRG 112
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 94.4 | 7.6e-30 | 194 | 306 | 1 | 91 |
EEEE-..-HHHHTT-EE--HHH.HTT.............---..............--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.............ggkke............esktltledesgrsWevkliyrkksgryvltkGWkeFvkang 73
f+k+ tpsdv+k++rlv+pk++ae+h + +++ l +ed g++W+++++y+++s++yvltkGW++Fvk++
Aco006706.1 194 FEKAVTPSDVGKLNRLVIPKQHAEKHfplqvvgsaspptA---TvaaaaamatatcKGVLLNFEDAAGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKS 288
89***************************99987655550...0555555666667999*************************************** PP
--TT-EEEEEE-SSSEE. CS
B3 74 LkegDfvvFkldgrsefe 91
Lk+gD+++F+++ e++
Aco006706.1 289 LKAGDVITFHRSTGPEKQ 306
**********76544444 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 5.43E-17 | 64 | 120 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.0E-8 | 64 | 112 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 4.68E-24 | 64 | 120 | No hit | No description |
| SMART | SM00380 | 1.4E-27 | 65 | 126 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.126 | 65 | 120 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.3E-20 | 65 | 120 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-39 | 191 | 320 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.60E-27 | 192 | 300 | No hit | No description |
| SuperFamily | SSF101936 | 1.31E-28 | 192 | 311 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 2.2E-23 | 194 | 315 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.1E-27 | 194 | 304 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.858 | 194 | 314 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 377 aa Download sequence Send to blast |
MEGSCIEEEV DKRPSPLRPP SMPLERVGSG ASVVLDPEGG GGGGGGGGGG GGGHEAESRK 60 LPSSKYKGVV PQPNGRWGAQ IYEKHQRVWL GTFNAEAEAA RAYDVAAQRF RGRDAVTNFR 120 PLSDTDDGDG ASELLFLSSH SKAEIVDMLR KHTYRDELRQ SRRALGPAAA AAAAGQKACR 180 GGGNYGSGGR DHLFEKAVTP SDVGKLNRLV IPKQHAEKHF PLQVVGSASP PTATVAAAAA 240 MATATCKGVL LNFEDAAGKV WRFRYSYWNS SQSYVLTKGW SRFVKEKSLK AGDVITFHRS 300 TGPEKQLFID YKPRSASAAA AAAAAAAGGP AGSGRAALRR EPGRDPGRGG GARARPMRRR 360 EAEESCRHAD VIRADV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 1e-53 | 193 | 322 | 13 | 125 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
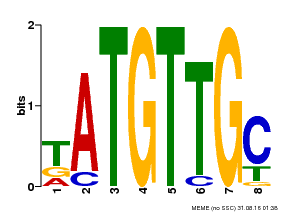
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM665315 | 0.0 | KM665315.1 Ananas bracteatus clone 50415d microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020083994.1 | 0.0 | LOW QUALITY PROTEIN: AP2/ERF and B3 domain-containing protein Os05g0549800-like | ||||
| Swissprot | Q6L4H4 | 1e-123 | Y5498_ORYSJ; AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| TrEMBL | A0A199V1A5 | 0.0 | A0A199V1A5_ANACO; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| STRING | XP_008791228.1 | 1e-143 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-106 | related to ABI3/VP1 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco006706.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



