 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco006046.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 296aa MW: 33688.1 Da PI: 6.8109 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 107.9 | 7.9e-34 | 18 | 110 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
F+ k+y++++d++++ li+w ++nsf+vl++ ef++ +Lp yFkh+nf+SFvRQLn+YgF+kv+ ++ weF+h+sF +g+++ll i
Aco006046.1 18 FVVKTYQMVNDPRTDVLIRWGMRNNSFLVLNPAEFSQFLLPCYFKHNNFSSFVRQLNTYGFRKVDPDR---------WEFAHESFLRGQTHLLPLIL 105
9********************999*****************************************999.........******************** PP
XXXXX CS
HSF_DNA-bind 99 rkkse 103
r+k++
Aco006046.1 106 RRKKK 110
99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.9E-36 | 12 | 101 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 9.5E-52 | 14 | 107 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 5.17E-33 | 16 | 107 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 1.9E-28 | 18 | 107 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-17 | 18 | 41 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-17 | 56 | 68 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 57 | 81 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-17 | 69 | 81 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 296 aa Download sequence Send to blast |
MEEKPVAYDG HEGHAAPFVV KTYQMVNDPR TDVLIRWGMR NNSFLVLNPA EFSQFLLPCY 60 FKHNNFSSFV RQLNTYGFRK VDPDRWEFAH ESFLRGQTHL LPLILRRKKK KSLSNEQSSS 120 NSRAKDSVVD EDEDEDVLVK EVTRLRQEQR LLEEELQGMN KRIQATERRP QQMMLFLAKA 180 ADEPESLSRL ILSKKQQFAA EKRRRLVNSP ITSTSTTTAP LTLTGPLMEL GFNEEVITTH 240 SVDEEKKPLY TVSSSVAGIE PSGWTEFGLE PEPKSASATD SVPFPFSLLG RGFFS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 1e-22 | 13 | 107 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 1e-22 | 13 | 107 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 1e-22 | 13 | 107 | 19 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 105 | 110 | RRKKKK |
| 2 | 201 | 205 | KRRRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
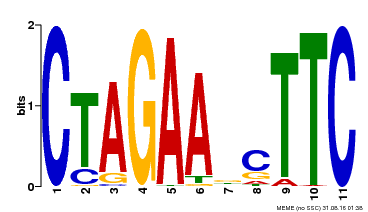
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020105591.1 | 0.0 | heat stress transcription factor C-1b-like | ||||
| Swissprot | Q6VBA4 | 4e-75 | HFC1A_ORYSJ; Heat stress transcription factor C-1a | ||||
| Swissprot | Q942D6 | 3e-76 | HFC1B_ORYSJ; Heat stress transcription factor C-1b | ||||
| TrEMBL | A0A199UDT8 | 0.0 | A0A199UDT8_ANACO; Heat stress transcription factor C-1b (Fragment) | ||||
| STRING | XP_008810034.1 | 2e-91 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11298 | 31 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 1e-43 | heat shock transcription factor C1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco006046.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



