 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Achn370001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; Ericales; Actinidiaceae; Actinidia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 574aa MW: 62418.1 Da PI: 4.6709 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.2 | 1.7e-17 | 37 | 84 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+v+++G g+W+++ ++ g+ R++k+c++rw ++l
Achn370001 37 KGPWTSAEDTILVDYVRKHGEGNWNAVQKHSGLFRCGKSCRLRWANHL 84
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 47.2 | 5e-15 | 90 | 133 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g+ T+eE+ l++++++ +G++ W++ a++++ gRt++++k++w++
Achn370001 90 KGAITPEEERLIIELHATMGNK-WARMAAHLP-GRTDNEIKNYWNT 133
6888******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.418 | 32 | 84 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.46E-29 | 36 | 131 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-14 | 36 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.3E-15 | 37 | 84 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-23 | 38 | 91 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.55E-11 | 39 | 84 | No hit | No description |
| PROSITE profile | PS51294 | 24.983 | 85 | 139 | IPR017930 | Myb domain |
| SMART | SM00717 | 4.4E-16 | 89 | 137 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.7E-14 | 90 | 133 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.4E-25 | 92 | 138 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.27E-11 | 94 | 133 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 574 aa Download sequence Send to blast |
MTNESDNEMI SKDCTGSPLM DEGNCDGSAS GGVVLKKGPW TSAEDTILVD YVRKHGEGNW 60 NAVQKHSGLF RCGKSCRLRW ANHLRPNLKK GAITPEEERL IIELHATMGN KWARMAAHLP 120 GRTDNEIKNY WNTRIKRRQR AGLPLYPPEV CLQALQENEQ NRNSNGDRGQ HDLLQANSYE 180 IPDVIFDSLK TNQGVLPYVP ELPDISASSI LLKGLGSSQY YGFVPPVVHR QKRLRESATS 240 YSGYDGSTRN EPPPFDHIPD DSCDNITTQS FGLSFPCDPD PITKNLLPFG GNQGSHSLLN 300 GNFSASKPAT PVAVKLELPS LQYPETDFSS WGSSSPPALL ESVDAFIQSP PPSGTDCLSP 360 RNSGLLDALL HEAKALSAKN HVSDKTSNSS TTSHTPGDVT HGSSLDTSKT KWEEYSESIS 420 PLGNSAASIF SERTPVSVSE NSLDEYPPAE TFPESCITFD LIVGSNMMLE AVEQKVWTLD 480 GEKNETATQL DFSRPDALLD SCWLGQSVTY ANDQEGVTTD AAAEAAMALG DDDSRNNDYK 540 PHMGAGTALN QGWGFGSCTW NNMPSVCQIS ELP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-31 | 35 | 138 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
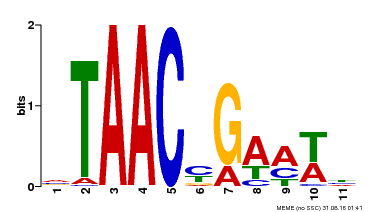
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028099441.1 | 0.0 | transcription factor GAMYB-like | ||||
| TrEMBL | A0A2R6R1D9 | 0.0 | A0A2R6R1D9_ACTCH; Transcription factor GAMYB like | ||||
| STRING | XP_008231582.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA14059 | 7 | 13 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-118 | myb domain protein 33 | ||||



