 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Achn195011 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; Ericales; Actinidiaceae; Actinidia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1024aa MW: 113840 Da PI: 6.201 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132 | 2e-41 | 156 | 233 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+ +lv+++ qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+q+
Achn195011 156 VCQVEDCGADLSNAKDYHRRHKVCEMHSKASRALVANVLQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNKRRRKTQP 233
6**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.3E-33 | 151 | 218 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.325 | 154 | 231 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.7E-38 | 155 | 236 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.9E-30 | 157 | 230 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 7.5E-6 | 308 | 327 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 7.5E-6 | 651 | 652 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 6.5E-7 | 789 | 905 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 7.5E-6 | 794 | 903 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.51E-6 | 794 | 904 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1024 aa Download sequence Send to blast |
METRVGGEAR LFYGMGMGSV DLRAVGKRSL EWDPNDWRWD GDLFIANPLN SNQSNYQSRQ 60 FLPIGEAGIP ITGGSSNSSS SCSDEVNLGT DKGKRELEKR RRVVVVEDDN LEDERSNLTL 120 KLGGHGHQIN QREVRNWDGT GGKKTKLLGT SSNRAVCQVE DCGADLSNAK DYHRRHKVCE 180 MHSKASRALV ANVLQRFCQQ CSRFHALQEF DEGKRSCRRR LAGHNKRRRK TQPDAVANSS 240 SLNDDQASGY LLISLLRILT NMHSNKSDET NDQDLLSHLL RSLASHGGLQ GDKNIYGLLQ 300 ESQNLLNSGS STGNSEMVSA LLSNSPQGLS KPIQHNLPVT TSEASQRGLC LENARTQVMQ 360 TTTSQNPSTI LSIKDNASPY SEARDISGGS RLNNFDLNDA YVDSDDCINL ERSPVPLGLG 420 TVSLECPSWV QQNSHQSSPP QTSRNSDSAS TQSPSSSSGE TQSRTDRIVL KLFGKEPSDF 480 PFVLRAQILD WLSHSPTDIE SYIRPGCVVL TIYLRLYDSA WEELRCDLTS SLSRLLDASD 540 DDTFWTTGWL YTRMQHQIAF MYNGQVVAET SLPLKNDTYS TIQSIKPIAV STSEGAQFLV 600 KGFNLKRPST RLLCALEGNY LVEEATEELL EGLDSPEGHD ELHFRNFSCS IPAVTGRGFI 660 EVEDNGLSSS FFPFIVAEKD VCSEIRTLER LIELPETDDI QGGNAKIEAR NRALDFVHEM 720 GWLLHRNHLK SRLGPLDPNS GLFPLRRFKW LMEFSVDHDW CAVVKKLLNI LLDGTVGAGE 780 QPFLKPALSE MGLLHRAVRR NSRSLVECLL KYVPENVSDE LRTEFSSLVG GDENFLFRPD 840 VVGPAGLTPL HVAAGRDGCE DVLDALTDDP GKVGVEAWKS ARDSTGFTPE DYARLRGHYS 900 YIHLVQKKIN RLAVTGHVVV DIPAYVPDCN INQKQHEQQQ QQQPASFQIG NSETRPIQRP 960 CRLCDQKLAY GNANSYLLYR PAMLSMVAIA AVCVCVALLF KSSPEVLYVF RPFRWELLDY 1020 GSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-31 | 151 | 230 | 5 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 98 | 102 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
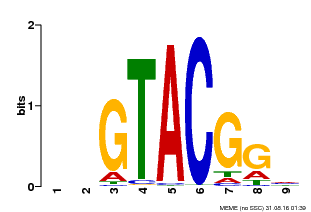
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028054458.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2R6PCT3 | 0.0 | A0A2R6PCT3_ACTCH; Squamosa promoter-binding-like protein | ||||
| STRING | VIT_07s0005g02260.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



