 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA9G00043 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | ZF-HD | ||||||||
| Protein Properties | Length: 325aa MW: 35687.6 Da PI: 8.4196 | ||||||||
| Description | ZF-HD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | ZF-HD_dimer | 107.7 | 6.5e-34 | 53 | 109 | 4 | 59 |
ZF-HD_dimer 4 vrYkeClkNhAaslGghavDGCgEfmpsegee.gtaaalkCaACgCHRnFHRrevee 59
++YkeClkNhAa+lGgha+DGCgEfmps+++ +++++lkCaACgCHRnFHRr++++
AA9G00043 53 FTYKECLKNHAAALGGHALDGCGEFMPSPSSIsSDPTSLKCAACGCHRNFHRRDPDD 109
68**************************9999899******************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04770 | 6.6E-28 | 54 | 107 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| TIGRFAMs | TIGR01566 | 1.8E-31 | 54 | 106 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| ProDom | PD125774 | 3.0E-14 | 55 | 119 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| PROSITE profile | PS51523 | 27.629 | 55 | 106 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| SuperFamily | SSF46689 | 2.11E-17 | 190 | 261 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.1E-29 | 190 | 261 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01565 | 3.0E-25 | 200 | 257 | IPR006455 | Homeodomain, ZF-HD class |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 325 aa Download sequence Send to blast |
MDMMPTSTPT PTTTTTPKSP EPESETPTRI QPAKPISFSN GIIKRHHHHP LLFTYKECLK 60 NHAAALGGHA LDGCGEFMPS PSSISSDPTS LKCAACGCHR NFHRRDPDDS TPSSAIPPPL 120 TAIEYHPHHR HHPPPPPPPP GHSRSPNSSS PPPISSSYML LALSGTNNNA GNNNLPFSDL 180 NFSGNNLSTH HHHHHAPGSR KRFRTKFSQF QKDKMQEFAE RVGWKMQKRD EDDIRDFCRQ 240 IGVDRGVLKV WMHNNKNTFN RRENLFSVGG TEIPKIAGDD TTNDDYNRDE ASGGGGGRGL 300 GHDLHQSVSS GGGGGFDKVK LFSLE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wh7_A | 9e-20 | 199 | 258 | 16 | 75 | ZF-HD homeobox family protein |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Putative transcription factor. Probably involved in establishing polarity during leaf development through the gibberellic acid (GA) signaling pathway. {ECO:0000269|PubMed:17387478}. | |||||
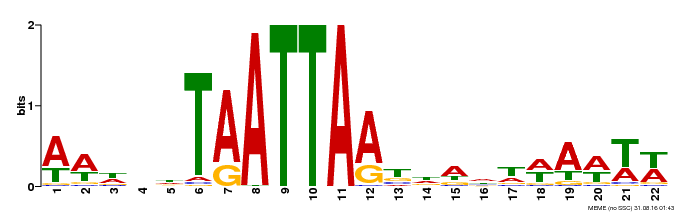
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00536 | DAP | Transfer from AT5G39760 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA9G00043 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellic acid (GA). {ECO:0000269|PubMed:17387478}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB016876 | 3e-86 | AB016876.1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MKM21. | |||
| GenBank | AK228484 | 3e-86 | AK228484.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-09-C01. | |||
| GenBank | AY086395 | 3e-86 | AY086395.1 Arabidopsis thaliana clone 249321 mRNA, complete sequence. | |||
| GenBank | AY091034 | 3e-86 | AY091034.1 Arabidopsis thaliana unknown protein (At5g39760) mRNA, complete cds. | |||
| GenBank | AY117347 | 3e-86 | AY117347.1 Arabidopsis thaliana unknown protein (At5g39760) mRNA, complete cds. | |||
| GenBank | CP002688 | 3e-86 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018458751.1 | 1e-112 | PREDICTED: zinc-finger homeodomain protein 9-like | ||||
| Swissprot | Q9FIW9 | 2e-98 | ZHD10_ARATH; Zinc-finger homeodomain protein 10 | ||||
| TrEMBL | A0A1J3G0A2 | 1e-122 | A0A1J3G0A2_NOCCA; Zinc-finger homeodomain protein 9 | ||||
| STRING | XP_006395326.1 | 1e-111 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2220 | 25 | 75 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G39760.1 | 5e-80 | homeobox protein 23 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



