 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA8G00350 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 554aa MW: 60960.8 Da PI: 7.9014 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 47.1 | 6e-15 | 224 | 273 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+++GV++++++grW+++I+d + k+++lg f+ta Aa+ +++a+ k++g
AA8G00350 224 SQFRGVTFYRRTGRWESHIWD------SgKQVYLGGFDTAHAAARVYDRAAVKFRG 273
789******************......44**********************99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 9.81E-15 | 224 | 283 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.17E-22 | 224 | 283 | No hit | No description |
| Pfam | PF00847 | 5.5E-7 | 225 | 273 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.9E-30 | 225 | 287 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 16.924 | 225 | 281 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.0E-15 | 225 | 281 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 554 aa Download sequence Send to blast |
MNKLVICFPF YGVFSMPLAT VYGSSTLQYS SGVDIFLAEL LAEHLSSVIS YFSFVIEHSL 60 VAAQRNESPD LFIDFRSLSI GSLEYELGSL LVLNRKAIVN FIGRLSVDLM DEPVTSNSSV 120 INADASSCID GEDSCSTPAV KFQFEILKGR CGEEDRDVVM TMEFFPVAEG DGNGMKYLDS 180 SAQSSKNFVD ICFGGGTGGN DAQVMQPPSQ PVKKSRRGPR SKSSQFRGVT FYRRTGRWES 240 HIWDSGKQVY LGGFDTAHAA ARVYDRAAVK FRGVEADINF NISDYEEDVK QLANLTKEEV 300 VQVLRRQGTA FSRSNSKYQG IALQKFGGWG AQMQQFHGNM GCDKAAIKWH GREGTLIEPH 360 ASRMVPEAAH VKLDLNLGIS LSLGDGPKQK DGSLRLHHGP NGIVCGRNYK SQIESHMAAA 420 PCDPPFNFLK RGPDHLINRH ILPSAFFSSK FNSSLQEIKP DKGLMSHSPR SFRTWQAQDH 480 PNGGTSATAT PSLLLSNAAS SRFSLPPMRP PSSITSLHLS QPFNLNLPGL HFTHPSDTTS 540 HHHHHPMNRP EAPP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Regulates negatively the transition to flowering time and confers flowering time delay. {ECO:0000250, ECO:0000269|PubMed:14555699}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00616 | PBM | Transfer from AT5G60120 | Download |
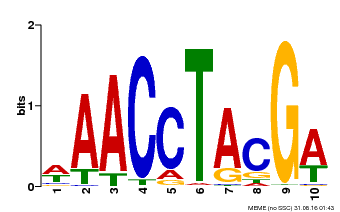
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA8G00350 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by miR172a-2/EAT. {ECO:0000269|PubMed:14555699}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006400879.1 | 0.0 | AP2-like ethylene-responsive transcription factor TOE2 | ||||
| Swissprot | Q9LVG2 | 0.0 | TOE2_ARATH; AP2-like ethylene-responsive transcription factor TOE2 | ||||
| TrEMBL | E4MWP9 | 0.0 | E4MWP9_EUTHA; mRNA, clone: RTFL01-19-B14 | ||||
| TrEMBL | V4LIS4 | 0.0 | V4LIS4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006400879.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM12836 | 17 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G60120.1 | 0.0 | target of early activation tagged (EAT) 2 | ||||



