 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA87G00209 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 820aa MW: 94806.6 Da PI: 7.2581 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 90.4 | 2.4e-28 | 63 | 150 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e++++ ++r++++++t+Cka+++vk+++dgkw +++++++HnHel p
AA87G00209 63 SFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESESS---NSRRSSVKKTDCKASMHVKRKSDGKWIIHEFVKDHNHELLP 150
5***************************************9998877...889999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 4.6E-26 | 63 | 150 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.0E-25 | 270 | 363 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.803 | 550 | 586 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 4.3E-5 | 552 | 584 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.9E-5 | 561 | 588 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 820 aa Download sequence Send to blast |
MDSQEHQVSI GGDANMSVVV VESHNRDVGS VDDLRSGDLG FSGDLDLEPR NGIDFESHEA 60 AYSFYQEYAK SMGFTTSIKN SRRSKKTKEF IDAKFACSRY GVTPESESSN SRRSSVKKTD 120 CKASMHVKRK SDGKWIIHEF VKDHNHELLP ALAYHFRIQR NVKLAEKNNI DILHAVSERT 180 RKMYVEMSRQ SGGYKNLGLP QTDVNTQLDK NRYLALEEGD SQVIFEYFKR IKKESPKFFY 240 AVDLNEEQRL RNIFWADAKS REDYMSFNDV VSFDTTYVKF NDKLPLALFI GVNNHCQPML 300 LGCALVADES KETFIWLIKT WLRAMGGKAP KVILTDQDKL LMNAISELLP NTRHCFALWH 360 VLEKIPEYFS HVMKRHENFL PKFNKCIFRS WADDQFEMRW WKLISRFGLQ DDEWLLWLHE 420 HRQKWVPSFM SDVFLAGMST SQRSESMNSF FDKYIHKKIT LKEFLRQYGA ILQNRYEEES 480 VADFDTCHKQ PALKSPSPWE KQMATTYTHT IFKKFQVEVL GVVACHPRKE KEDGNIVTFR 540 VQDCEKDDEF LVTWSKTKSE LCCFCHMFEY KGFLCRHALM ILQICGFASI PPSYILKRWT 600 KDAKTGLVAS EEADQVQTRV QRYNDLCHRA IELSEEGCVS EENYNIVLRT LVESLKNCVD 660 LNNSRNCITE YSSQINGVHE EDNQVMAPVK ATKKKTVVRK RKGEPEVSQM LESQQSLQSM 720 ENLSSEGISI NGYYGPQQNV QGLLNLMEPP HEGYYVNQRT IQGLGQLNSI APVQDSFFAN 780 QQAMPGLGQI DFRPPPNFTY NLQDEHLRSA QLPGSSSRHL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
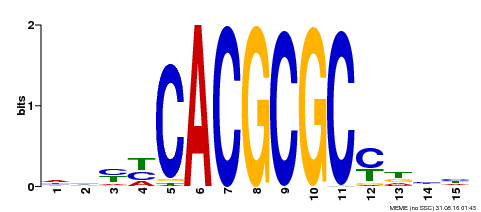
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA87G00209 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006414566.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A1J3JL92 | 0.0 | A0A1J3JL92_NOCCA; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| STRING | XP_006414566.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



