 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA54G00241 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 934aa MW: 102835 Da PI: 5.6151 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.9 | 4.8e-29 | 577 | 627 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
AA54G00241 577 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 627
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.757 | 566 | 647 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.1E-26 | 579 | 627 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 7.2E-23 | 835 | 923 | No hit | No description |
| Pfam | PF00564 | 2.3E-19 | 838 | 919 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 26.856 | 838 | 920 | IPR000270 | PB1 domain |
| SMART | SM00666 | 6.5E-28 | 838 | 920 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 3.3E-27 | 838 | 920 | No hit | No description |
| CDD | cd06407 | 1.12E-33 | 839 | 919 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 934 aa Download sequence Send to blast |
MCEPEDDTTR NGVITSVSRS REIMMDVDDL DLDGSWPLDQ ISYLSSANRM MSPMFVTSSS 60 EQPCSPLWAF SDGGGNGIHG NGGGDDEKRS AASVVPSFRF AEYPLFLPLS NSLKSLEIAD 120 SSPSVAESNA EKNNHIQFPS PLMGLVPPPN TDNYCVWAPV RKNGRNLLTT LGQPFVLNPN 180 GNGLNQYRMI SLTYMFSVDS ESDIELGLPG RVFRQKLPEW TPNVQYYSSK EFSRLDHALH 240 YNVRGTLALP VFHPSGQSCI GVVELIMTSE KIHYAPEVDR VCKALEAVNL KSSEILDHQP 300 AQICNESRQN ALAEILEVLT VVCETHNLPL AQTWVPCQHG SVLANGGGLK KNCTSFDGSC 360 MGQICMSTTD MACYVVDFHV WGFRDACLEH HLQKGQGVAG RAFLNGGSCF CRDITKFCKT 420 QYPLVHYALM FKLTTCFAIS LQSSYTGDDS YILEFFLPST ITDEHEQDSL LGSLLVTMKE 480 HFQSLRVASG VDFGEDDDKL SFEIIQALPD QKINSKIESI RVPFSGFKAN AREPPLNPQP 540 AVLTSDPVNE KSNNVATTVN GVAKEKKKTE KKRGKTEKTI SLDVLQQYFT GSLKDAAKSL 600 GVCPTTMKRI CRQHGISRWP SRKIKKVNRS ITKLKRVIES VQGTDGGLDL TTMAVGSIPW 660 THSQTPPQPL NSPNGHKPSE LPNTNNSPNH WSSDHSPQEP NGSPELPSNG YKSAGTPTSH 720 GSCDGNQLEE TKTLNQDPLF TVGGSPGLMC PPYSRELDVS ATSFSLQNRL LGSIDHFRGM 780 PIADVGSSKD LRNLCSSTAF EDKFPDSNWM SNENSSNNIY VPALEEPLAT TRDPNTRTVT 840 IKASYKEDII RFRISSGSGL MELKDEVAKR LKLDAGIFDI KYLDDDNEWV LIACDADLQE 900 CLEIPRLSRT NIVRLLVHDV TTNLGSSCES TGEL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
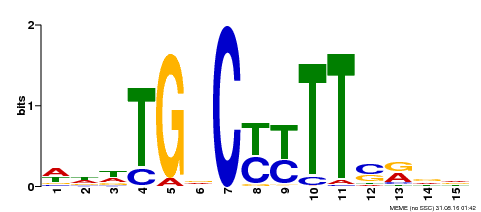
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA54G00241 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006413484.1 | 0.0 | protein NLP7 | ||||
| Refseq | XP_024005488.1 | 0.0 | protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | V4MHY8 | 0.0 | V4MHY8_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006413484.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



