 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA44G00309 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 565aa MW: 63181.1 Da PI: 6.7572 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 31.5 | 4.4e-10 | 259 | 315 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng...krkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + rk + g ++ +++Aa+a++ a++k+++
AA44G00309 259 SIYRGVTRHRWTGRYEAHLWDnSCR-RegqARKGRQ-GGYDKEDKAARAYDLAALKYWN 315
57*******************4444.2344336655.7799999*************97 PP
| |||||||
| 2 | AP2 | 49.7 | 9.2e-16 | 358 | 409 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
AA44G00309 358 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 409
57****************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.03E-13 | 259 | 324 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 7.4E-8 | 259 | 315 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.42E-17 | 259 | 325 | No hit | No description |
| PROSITE profile | PS51032 | 16.753 | 260 | 323 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.7E-10 | 260 | 323 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.1E-17 | 260 | 329 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.0E-6 | 261 | 272 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.58E-11 | 358 | 417 | No hit | No description |
| SuperFamily | SSF54171 | 1.05E-17 | 358 | 418 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 3.9E-10 | 358 | 409 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.111 | 359 | 417 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.2E-18 | 359 | 417 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.9E-31 | 359 | 423 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.0E-6 | 399 | 419 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 565 aa Download sequence Send to blast |
MAPLTNWLTF SLSPMEMLRS SDQSHFVSYD ASSPVSSPYL LDNFYGWTNQ KPQEFFKEEA 60 QIAASMADST ILSTYVDSQT HHIPKLEDFL GDSSSIVRYS DNSQTETQDS SLTQIYDPRH 120 HQNHNHHSTT AGFYSDHHDF KAMANFQTAF STNSGSEVDD STSIGRNHLT GTGGEYLGHV 180 VESSGSELGF HGGSSTGGAL SLAVNNNNNT NHITDPYNTE ERINNNNDKI DSEKEKPIVA 240 VETSDSSKKI ADTFGQRTSI YRGVTRHRWT GRYEAHLWDN SCRREGQARK GRQGGYDKED 300 KAARAYDLAA LKYWNATATT NFPIANYSKE LEEMKHMTKQ EFIASLRRKS SGFSRGASIY 360 RGVTRHHQQG RWQARIGRVA GNKDLYLGTF ATEEEAAEAY DIAAIKFRGI NAVTNFEMNR 420 YDVEAIMKSA LPIGGAAKRL KLSLESSSEQ KPILGQHNHH HLQQQQQQQI QSSPNHSINF 480 ALCPNSGVQS QIPCGIPFEA AALYHHQQQQ QQQQQQQQQN FFQHFPVTAG SDSTGSNSTV 540 QASMGLMAPN AAEFFLWPNH NQSNY |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
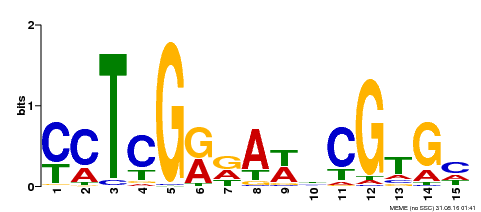
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA44G00309 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006399549.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Refseq | XP_024011965.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A1J3E758 | 0.0 | A0A1J3E758_NOCCA; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| STRING | XP_006399549.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3333 | 27 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



