 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA32G00026 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 794aa MW: 87198.6 Da PI: 6.5064 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 63.5 | 3.1e-20 | 90 | 145 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++ +p++++r eL+k+l L++rqVk+WFqNrR+++k
AA32G00026 90 KKRYHRHTPQQIQELESMFKECAHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 145
688999***********************************************999 PP
| |||||||
| 2 | START | 194.1 | 6.6e-61 | 293 | 519 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE......EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECT CS
START 1 elaeeaaqelvkkalaeepgWvkss......esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....kaetleviss 87
ela++a++elvk+a +ep+Wvks+ +++n++e+++ f+++k+ + +ea+r+sg+v+ ++ lve+l+++ +W+e+++ +a+t++vis
AA32G00026 293 ELALTAMDELVKLAHTDEPLWVKSVeggrerDELNQEEYTRAFSSNKStgFVTEASRTSGMVIINSLALVETLMNSD-RWTEMFPcnvaRATTTDVISA 390
5899****************************************9999999**************************.********************* PP
T......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSX CS
START 88 g......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrl 176
g gal+lm+aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d + ++ s+s ++lpSg+++++++ g+skvtwveh+++++++
AA32G00026 391 GmagtrnGALHLMNAELQVLSPLVPvRNVNFLRFCKQHAEGVWAVVDVSIDAAIPENNsgpSQSHLSIRRLPSGCVVQDMPTGYSKVTWVEHAEYEESQ 489
**************************************************9765544444644444445****************************** PP
XHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 177 phwllrslvksglaegaktwvatlqrqcek 206
+h+l+r+l++sgl +g ++w+atlqrqce+
AA32G00026 490 IHELYRPLLRSGLGFGSQRWLATLQRQCEC 519
****************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.13E-20 | 77 | 147 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-21 | 79 | 147 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.941 | 87 | 147 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.5E-17 | 88 | 151 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.07E-17 | 89 | 147 | No hit | No description |
| Pfam | PF00046 | 1.0E-17 | 90 | 145 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 122 | 145 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.598 | 284 | 522 | IPR002913 | START domain |
| CDD | cd08875 | 1.94E-111 | 288 | 518 | No hit | No description |
| SuperFamily | SSF55961 | 8.93E-28 | 288 | 519 | No hit | No description |
| Pfam | PF01852 | 1.0E-53 | 293 | 519 | IPR002913 | START domain |
| SMART | SM00234 | 1.6E-36 | 293 | 519 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.19E-21 | 553 | 778 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 794 aa Download sequence Send to blast |
MAQAAAAVTL FSPPLTKSAF ASSGLSLALE QPERGTNVGE TSMRNGGDNF DGISVNRRSR 60 EEEHESRSGS DNLEGISGED HDATDKPPRK KRYHRHTPQQ IQELESMFKE CAHPDEKQRL 120 ELSKRLCLET RQVKFWFQNR RTQMKTQLER HENALLRQEN DKLRAENMSI REAMRNPICT 180 NCGGPAILGD VSLEEHHLRI ENARLKDELD RVCAITGKFL GRHITDHNHH TLPPNSSLEL 240 AVGSNGTGGF AFPPDFGGGC LSVVPTPPPH QQQQQQQHPT GINGIDQRSV LLELALTAMD 300 ELVKLAHTDE PLWVKSVEGG RERDELNQEE YTRAFSSNKS TGFVTEASRT SGMVIINSLA 360 LVETLMNSDR WTEMFPCNVA RATTTDVISA GMAGTRNGAL HLMNAELQVL SPLVPVRNVN 420 FLRFCKQHAE GVWAVVDVSI DAAIPENNSG PSQSHLSIRR LPSGCVVQDM PTGYSKVTWV 480 EHAEYEESQI HELYRPLLRS GLGFGSQRWL ATLQRQCECL TILMSSSATS HDNSLTLTYT 540 IYICTQWKAA ITPGGRKSML RLAQRMTFNF CCGISAPSVH SWSKLSVGNV DRDVRVMTRK 600 SVDDPGEPPG IVLSAATSVW LPASPQRLFD FLRNERLRCE WDILSNGGPM QEMAHIAKGQ 660 DHGNSVSLLR SNALNANQSS MLILQETCID ASGALVVYAP VDIPAMHVVM NGGDSSCVAL 720 LPSGFAVLPD GGGPEGTSEH TQRPLGGGSL LTVAFQILVN NLPTAKLTVE SVETVNNLIS 780 CTVQKIRAAL QCDT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
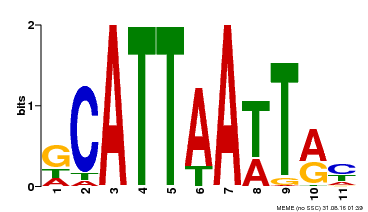
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA32G00026 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353253 | 0.0 | AK353253.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-28-B24. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020878511.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A1J3DE85 | 0.0 | A0A1J3DE85_NOCCA; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| STRING | Bostr.10064s0036.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



