 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA29G00253 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 414aa MW: 47448 Da PI: 6.6998 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.7 | 9.4e-32 | 84 | 137 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH +F++ave+LGG+++AtPk +l+lm+vkgL+++hvkSHLQ+YR+
AA29G00253 84 PRLRWTPELHLCFLQAVERLGGPDRATPKLVLQLMNVKGLSIAHVKSHLQMYRS 137
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-30 | 80 | 138 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.321 | 80 | 140 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.94E-16 | 81 | 139 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.7E-24 | 84 | 139 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.0E-9 | 85 | 136 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 414 aa Download sequence Send to blast |
MRSNTKSSEN SKTTCPSKEE DKDEDDEEEE DDEEDQSPST NNSYVEESGS HDHHDDDDHD 60 HHHDHNQIKK NGGSVRPYNR SKTPRLRWTP ELHLCFLQAV ERLGGPDRAT PKLVLQLMNV 120 KGLSIAHVKS HLQMYRSKKT DDPNQGDQGF SFEQGSTYTY NLSQLPMLQS FDQRPSSSLG 180 YGGSSWTDHR RQIYRSPWRG LTARDNTRTR QTLFSSSQPI ERFHGVSNNT ILDDHKNKTI 240 SFRLDSHQAA ANENEAETVP RIHRSFLQGM KTFNKSLGQS FSNPCSSTTT RPIQDHIATT 300 LRSSYPRDSI RNIEESSENV LKRKRLLFTN EINKTDQDLD LSLSLKVSQR YEKIKECMLE 360 DEEQEDNNNN GLSLSLSTSC LTKLGRSIRK EDNDQKDHKK RKISVLASPL DLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 1e-17 | 85 | 139 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-17 | 85 | 139 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-17 | 85 | 139 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-17 | 85 | 139 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-17 | 85 | 139 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 388 | 400 | RKEDNDQKDHKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
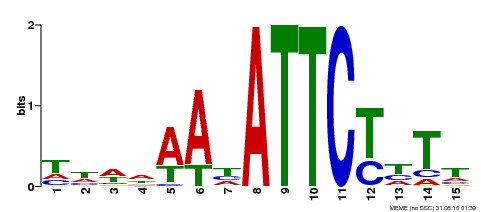
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA29G00253 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002881704.1 | 1e-176 | uncharacterized protein LOC9315925 | ||||
| Refseq | XP_013634613.1 | 1e-177 | PREDICTED: uncharacterized protein LOC106340283 isoform X2 | ||||
| TrEMBL | A0A1J3CY76 | 0.0 | A0A1J3CY76_NOCCA; Putative Myb family transcription factor | ||||
| TrEMBL | A0A1J3GUA0 | 0.0 | A0A1J3GUA0_NOCCA; Putative Myb family transcription factor | ||||
| STRING | Bo4g025360.1 | 1e-176 | (Brassica oleracea) | ||||
| STRING | fgenesh1_pm.C_scaffold_4001775 | 1e-176 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9359 | 21 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 1e-166 | G2-like family protein | ||||



