 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA17G00028 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 844aa MW: 97522 Da PI: 7.6233 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 81 | 2.1e-25 | 91 | 193 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtkleleHnH 87
+fY+eY++ +GF++ +++s++sk+++e+++++f Cs++g+++e +k+ e+ +r+ +t+Cka+++vk++ dgkw++++++ eHnH
AA17G00028 91 SFYQEYSRAIGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDKSfnrprarqskqdPENMAGRRTCAKTDCKASMHVKRRPDGKWVIHSFVREHNH 189
5*******************************************99999999877654444444499999***************************** PP
FAR1 88 elap 91
el p
AA17G00028 190 ELLP 193
*975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 2.9E-23 | 91 | 193 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.6E-25 | 291 | 383 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.359 | 571 | 607 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.7E-7 | 582 | 609 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 844 aa Download sequence Send to blast |
MDINLRLPSG DLCKEDDEER GIDNVLQIEH KLHEAEEDMD IGKIEDGTVE LHTDDSAGMS 60 VPGGELVLYR EGMNLEPLNG MEFESHGEAY SFYQEYSRAI GFNTAIQNSR RSKTTREFID 120 AKFACSRYGT KREYDKSFNR PRARQSKQDP ENMAGRRTCA KTDCKASMHV KRRPDGKWVI 180 HSFVREHNHE LLPAQAVSEQ TRKIYAAMAK QFAEYKTVIS LKCDAKSSFE KGRNVSVETG 240 DFKILLDFLS RMQNLNSNFF YAVDLGEDQR VKNVFWVDAR SRYNYSSFCD VVSLDTTYVR 300 NKYKMPLAIF VGVNQHYQYM VLGCALISDE NAATFSWIME TWLKAMGGQA PKVLITEHDI 360 TMNSIVPEIM PNTRHCFFLW HVLMKVSENL GQVMKQHDNF MPKFEKCIYK SWKDEDFARR 420 WWKILARFGL KEDQWMQSLY EDRRKWAPAY MTDVLLAGMS TSQRADSINA FFDKYMHKKT 480 SVQEFVKLYD TILQDRYEEE AKADSETWNK QPAMKSPSPF EKSVSEVYTP AVFKKFQSEV 540 LGAIACCPRE QNRDGTCSTF RVQDFENNQD FVVTWNQAKA EVSCICRLFE YKGYLCRHTL 600 NVLQSCHLSS IPSQYILKRW TKDAKGRPFP REPQQLQTRL QRYNDLCQRV LKLNEEASFS 660 QESYNIAFRA IEEALRNCAA VNTSGRNPQD VVNSPTQGLI SVEEDNQMRS KTSKKKNPTK 720 KRKVNSEQEV MPVAAPESLQ QMDKLSPRTV GIESYYGTQQ SVQGMVQLNL MGPTRDNFYG 780 NQQTIQGLRQ LNSIAPSYDS YYSAQQGIHG QGVDFFRPTN FSYDIRDDPN VRATQLHEDA 840 SRHS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
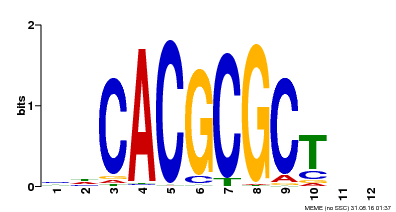
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA17G00028 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020886687.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A1J3CXP8 | 0.0 | A0A1J3CXP8_NOCCA; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| STRING | Bostr.19424s0443.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



