 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK44549.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 347aa MW: 39042.2 Da PI: 9.8064 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.9 | 1.1e-12 | 54 | 102 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+ ++ a++k++g
KFK44549.1 54 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEQEAAESYDIAARKFRG 102
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 98.3 | 4.8e-31 | 173 | 285 | 1 | 96 |
EEEE-..-HHHHTT-EE--HHH.HTT.................---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.................ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvF 82
f+k tpsdv+k++rlv+pk++ae+h ++++++ + led++g++W+++++y+++s++yvltkGW++Fvk+++L++gD v F
KFK44549.1 173 FDKSVTPSDVGKLNRLVIPKQHAERHfplpaftttetvtmtmtVASPTKGVLINLEDRTGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKNLRAGDLVCF 271
899******************************99999999996667899************************************************* PP
EE-SSSEE..EEEE CS
B3 83 kldgrsefelvvkv 96
+++ +++l++ +
KFK44549.1 272 ERSTGPDRQLFIDW 285
*8766777777765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 6.70E-25 | 54 | 109 | No hit | No description |
| Pfam | PF00847 | 2.9E-7 | 54 | 102 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.17E-16 | 54 | 110 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.1E-19 | 55 | 110 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.888 | 55 | 110 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.1E-28 | 55 | 116 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-39 | 170 | 289 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.98E-27 | 171 | 274 | No hit | No description |
| SuperFamily | SSF101936 | 2.49E-30 | 171 | 284 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 1.1E-27 | 173 | 285 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.493 | 173 | 289 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.1E-23 | 173 | 289 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 347 aa Download sequence Send to blast |
MDYSCVEDTS TTSESISPSP AAMRLYRMGS GGSSVVLDTD QNGVETESRK LPSSKYKGVV 60 PQPNGRWGAQ IYEKHQRVWL GTFNEEQEAA ESYDIAARKF RGRDAVINFK SLVRNDAESV 120 FLDTHSKSEI VDMLRKHTYK DELQQSKRKF LDNGKLKGNE NDVVLRGVRE FLFDKSVTPS 180 DVGKLNRLVI PKQHAERHFP LPAFTTTETV TMTMTVASPT KGVLINLEDR TGKVWRFRYS 240 YWNSSQSYVL TKGWSRFVKE KNLRAGDLVC FERSTGPDRQ LFIDWKVRSG TVQNVVRLFG 300 VNIYSDTKPN DVVEECGGGG GGKKRYRGGA DLLALGCSKK QAIINVL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 1e-54 | 170 | 291 | 11 | 120 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 321 | 327 | GKKRYRG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
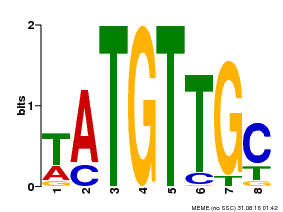
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK44549.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002890683.1 | 0.0 | AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| Swissprot | Q9C6M5 | 0.0 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A0A087HQZ7 | 0.0 | A0A087HQZ7_ARAAL; Ap2 erf and b3 domain-containing transcription factor rav2 | ||||
| STRING | A0A087HQZ7 | 0.0 | (Arabis alpina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 1e-178 | RAV family protein | ||||



