 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK28113.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1060aa MW: 118560 Da PI: 5.822 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.7 | 8.5e-57 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkD ++w+kkkdgktv+E+hekLKvg++++l+cyYah+e+n+ fqrrcy
KFK28113.1 19 LSEaQHRWLRPAEICEILRNHQKFHISSEPPNRPSSGSLFLFDRKVLRYFRKDAHNWRKKKDGKTVKEAHEKLKVGSIDMLHCYYAHGEDNEYFQRRCY 117
45559********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
w+Le++l +iv+vhylevk
KFK28113.1 118 WMLEQDLMHIVFVHYLEVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.125 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.2E-83 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.3E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 6.1E-4 | 464 | 554 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 2.4E-14 | 470 | 556 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 1.38E-19 | 648 | 763 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 20.473 | 652 | 734 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 5.5E-8 | 653 | 729 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.1E-18 | 655 | 767 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.17E-15 | 658 | 761 | No hit | No description |
| PROSITE profile | PS50088 | 9.351 | 669 | 701 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1500 | 669 | 698 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0014 | 702 | 731 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.247 | 702 | 734 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.19E-8 | 876 | 926 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.11 | 876 | 898 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.425 | 878 | 906 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0017 | 878 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 8.9E-4 | 899 | 921 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.706 | 900 | 924 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 3.7E-4 | 901 | 921 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1060 aa Download sequence Send to blast |
MADRGSFGFA PRLDIQQLLS EAQHRWLRPA EICEILRNHQ KFHISSEPPN RPSSGSLFLF 60 DRKVLRYFRK DAHNWRKKKD GKTVKEAHEK LKVGSIDMLH CYYAHGEDNE YFQRRCYWML 120 EQDLMHIVFV HYLEVKGNRT SSSGIKENNS NSLSGTASVN IDSAATPSST LSPLCEDADS 180 GDSHQASSSL QANREPQSVA PLIRHQQNAS TMNSYNPSSI LGNGDGWTSA PGIGIVSQVH 240 GNRVKDNESQ RSVDVPAWDA SFENSLARYQ NLPYNALLTQ TQHSNTGLMP VGGNTEKGSP 300 LMAEHLRNPL QTQLNWQIPV QDSLPLQKWP MDSHSGMTDD ADLALLEQRA RENFGTFSSL 360 LGSQNQQPIM GSFQAPFTSI EAAYIPQLGP EDLLYEASAN QTLPLRKSLL KKADSLKKVD 420 SFSRWVNNEL GEMEDLHMQS SSEGLAWTAV GTAAAASSLS PSLSEHQRFT IIDFWPKSSQ 480 TDEEVEVMVI GTFMLSPQDV TSYSWSCMFG EVEVPAEILV DGVLYCHAPP HEIGQVPFYV 540 TCSDRFSCSE VREFDFLPGS PKKFDTTDIY GAFTNETSLH LRFANLLARR SLVQEHIFED 600 VGEKRRKIRD IMLLIDEKES LVPGTTEKDL PEMEARERLI REEFEDKLYL WLIHMVTEEG 660 KGPNILDTEG QGVLHLAAAL GYDWAIKPIL AAGVSINFRD ANGWSALHWA AFNGREDTVA 720 VLVSLGADAG ALTDPSPELP LGKTASDLAY GNGHRGISGF LAESSLTSYL EKLTVDAKEN 780 SSADSSKAKA VQTVAERTAT PISYSDVPET LSMKDSLTAV LNATQAADRL HQVFRMQSFE 840 RKQLSEFGDD QLDISDELAV SFAAAKSKKA GHSNGTVHAA AVQIQKKYRG WKKRKEYLLL 900 RRRVVKIQAH VRGHQVRKQY RTIIWSVGLL EKIILRWRRK GSGLRGFKRE TVTMSPEPVS 960 VAPQEDDYDF LKEGRKQTEE RLQKALTRVK SMAQYPEARA QYRRLLTVVE GFRENEASSS 1020 SAMNNNNSNT EEAANYNEDD DLVDIDSLLD DDTFMSLAFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
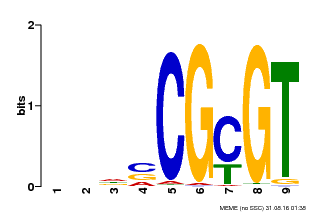
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK28113.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394184.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A087GE11 | 0.0 | A0A087GE11_ARAAL; Uncharacterized protein | ||||
| STRING | A0A087GE11 | 0.0 | (Arabis alpina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



