 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK25008.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 249aa MW: 28312.8 Da PI: 8.8844 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.2 | 7.9e-08 | 30 | 75 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqk 46
+WT+eE +++ +a + + + +W ++a++++ g+t ++ + k
KFK25008.1 30 SWTKEENKKFERALAVYADDtpdRWFKVASMIP-GKTISDVMRQYSK 75
7*****************99*************.******9998877 PP
| |||||||
| 2 | Myb_DNA-binding | 40.2 | 7.9e-13 | 132 | 176 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++G+g+W+ I+r + +t+ q+ s+ qky
KFK25008.1 132 PWTEEEHRRFLLGLLKYGKGDWRNISRNFVGSKTPTQVASHAQKY 176
8****************************999************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-4 | 25 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51293 | 8.348 | 26 | 79 | IPR017884 | SANT domain |
| SMART | SM00717 | 7.5E-8 | 27 | 79 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 4.09E-11 | 29 | 84 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.07E-7 | 30 | 76 | No hit | No description |
| Pfam | PF00249 | 9.7E-7 | 30 | 75 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 18.311 | 125 | 181 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-16 | 127 | 180 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 9.9E-11 | 129 | 179 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 1.2E-16 | 129 | 180 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-11 | 129 | 181 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 9.19E-10 | 132 | 176 | No hit | No description |
| Pfam | PF00249 | 3.4E-10 | 132 | 176 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 249 aa Download sequence Send to blast |
METLHPLLSH VPTSDHRFVV QEMMSLQSSS WTKEENKKFE RALAVYADDT PDRWFKVASM 60 IPGKTISDVM RQYSKLEEDL FDIEAGLIPI PGYNSATPCG FDELVSPRDF DLYRKRPNGA 120 RGLDQERRKG VPWTEEEHRR FLLGLLKYGK GDWRNISRNF VGSKTPTQVA SHAQKYYQRQ 180 LSGAKDKRRP SIHDITTVNL LNANLSRPDH GAESKLGFTD SDNAEKGVMF LGQHLSSVFK 240 STGANDYGF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 8e-16 | 23 | 95 | 1 | 75 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
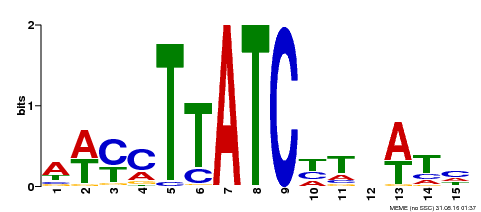
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00489 | DAP | Transfer from AT5G05790 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK25008.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY519531 | 0.0 | AY519531.1 Arabidopsis thaliana MYB transcription factor (At5g05790) mRNA, complete cds. | |||
| GenBank | BT026054 | 0.0 | BT026054.1 Arabidopsis thaliana At5g05790 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001332439.1 | 1e-167 | Duplicated homeodomain-like superfamily protein | ||||
| Refseq | NP_196198.1 | 1e-167 | Duplicated homeodomain-like superfamily protein | ||||
| Swissprot | Q8S9H7 | 9e-75 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A087G556 | 0.0 | A0A087G556_ARAAL; Uncharacterized protein | ||||
| STRING | A0A087G556 | 0.0 | (Arabis alpina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4340 | 26 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G05790.1 | 1e-171 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



