 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna15311.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 778aa MW: 85645.1 Da PI: 6.6805 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 85 | 5.7e-27 | 385 | 450 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf++l++++e+gi++ln++a++L +v ++RRiYDi+NVLe+++liekk kn+irwkg
mrna15311.1-v1.0-hybrid 385 RYDSSLGLLTKKFITLIKQAEDGILDLNKAAETL---EV--QKRRIYDITNVLEGIGLIEKKLKNRIRWKG 450
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.1E-28 | 382 | 452 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 2.31E-16 | 385 | 448 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 3.2E-34 | 385 | 450 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| Pfam | PF02319 | 4.3E-24 | 387 | 450 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF144074 | 2.3E-31 | 464 | 567 | No hit | No description |
| CDD | cd14660 | 3.39E-47 | 466 | 567 | No hit | No description |
| Pfam | PF16421 | 8.5E-31 | 466 | 565 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051446 | Biological Process | positive regulation of meiotic cell cycle | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 778 aa Download sequence Send to blast |
MAGLWRTMGT DDVAETRARS GSVVGDWIYP DLVAIPCTFV FPIGAPNPVT EGGDERSKAG 60 FYNQFGRLGF EGRFLTFWRL ADEIDSMAFG VQRRRRCGNL GGATMAVGLS VLHALWVGSE 120 LFRLRRFSDL FLVYNNSYLL LVEMGGLCPG NRTQRAMFVI IRRRVPCLVK WSQPPSGKMK 180 LGRSPSPSPL NPKPPPIPSA ALRFPQTLKI YPLLSPLVVF ANPRRSATVL AGRSTSMSGL 240 ARAPNRQPAP PPPPTGGGNG QIVPPMRRHL VFESNKPPFF PPADYHQFSG GGQRAADQEP 300 EAIVVRSPQF KRKGTSDNNE AENNDRTSSP GYNNISQSPF QTPVSTTKAG KTNNRTKASK 360 GKSGPQTPAS NAGSPCPLTP AGSCRYDSSL GLLTKKFITL IKQAEDGILD LNKAAETLEV 420 QKRRIYDITN VLEGIGLIEK KLKNRIRWKG FDASRPGDLD ADLAILQGEV ENLSLKERSL 480 DDRIREMEEK LRDLSGDENN RKWLFVTEED IKGLPCFQNE TLIAIKAPHG TTLEVPDPDE 540 AVDYPQRRYR IILRSTMGPI DVYLVSQFEE KFDDVNGAEP PVSFPIASVC GTDEQSTAEM 600 VAGHDRGKEI EPRVQQAQVD QTCSDVNASQ EFGGGMMKIV PSDVDNDADY WLLSGVDVSI 660 TDMWRTDSAI EWNGEDMLNA DFGLPDVSTP RPQTPPSGIV EVPSHAANYP QSFRIEYQES 720 HLDIDIDIPH PSGHDVELGH LNLLPLKNKN YRPGSQIACQ NLKILNYLVS ILILTLG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 6e-24 | 380 | 451 | 2 | 74 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Regulates gene expression both positively and negatively. Activates the expression of E2FB. Involved in the control of cell-cycle progression from G1 to S phase. Stimulates cell proliferation and delays differentiation. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11889041, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:15377755, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:19662336}. | |||||
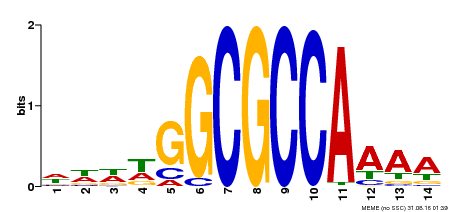
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00293 | DAP | Transfer from AT2G36010 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna15311.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004305631.1 | 0.0 | PREDICTED: transcription factor E2FA isoform X1 | ||||
| Swissprot | Q9FNY0 | 1e-169 | E2FA_ARATH; Transcription factor E2FA | ||||
| TrEMBL | A0A2P6RCD7 | 0.0 | A0A2P6RCD7_ROSCH; Putative transcription factor E2F-DP family | ||||
| STRING | XP_004305631.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1947 | 34 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G36010.1 | 1e-174 | E2F transcription factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna15311.1-v1.0-hybrid |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



