 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | model.Picochlorum_contig_68.g377.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Picochlorum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 648aa MW: 72304.6 Da PI: 6.3542 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 130.3 | 7.5e-41 | 38 | 153 | 4 | 116 |
CG-1 4 e.kkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLK 74
+ ++ wlknee++a+Le+f+ +l+ +++t+p+ g+l +++rk+ r fr+D ++w+kk d kt++E+hekLK
model.Picochlorum_contig_68.g377.t1 38 KaESSWLKNEEVLAVLEKFQGLDLSWpqQAATQPEGGRLYIFSRKVCRAFRNDKHTWRKKNDNKTIKETHEKLK 111
44899***************99986557999******************************************* PP
CG-1 75 vggvevlycyYahseenptfqrrcywlLeeelekivlvhyle 116
v+ e+l+cyYah+++n +qrrcyw+L+++++++v+vhyl
model.Picochlorum_contig_68.g377.t1 112 VQDKETLNCYYAHADRNDGLQRRCYWILDKRYDDVVMVHYLC 153
****************************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 50.999 | 32 | 160 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.8E-40 | 35 | 155 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 6.4E-38 | 39 | 152 | IPR005559 | CG-1 DNA-binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 648 aa Download sequence Send to blast |
MKDLVISRIP PGEEKRQAVP LDPEAAELGE NVMEIVKKAE SSWLKNEEVL AVLEKFQGLD 60 LSWPQQAATQ PEGGRLYIFS RKVCRAFRND KHTWRKKNDN KTIKETHEKL KVQDKETLNC 120 YYAHADRNDG LQRRCYWILD KRYDDVVMVH YLCTSTSRVA GRVSMGQAGS MTPSGSGLSA 180 GKSSRPKREA AMRKKRYSEI FGEDSVELAG TNLKREEPVQ EQRMKEAPRH DAREDLEPYD 240 SFLERIPSDM GVDPSLYPGD LEVSAAQIVR VADSSPRKSQ MGDDVDRKLS FGKYDLKALL 300 EGSTSPNKIS SMTEEEYLVR GFSKVDSLGY LRNFSFETKM GNGDVFDGHT GGAGVTFLHR 360 GNGSTELDKF ARSDSYNEWQ PSSSTDPESS GVRQNEKVET LRRFVIPGSD SSLSSFASQG 420 FDQSNNLRKR SNPTSTVPPK SMDVRMTEAA ILSQDANIRR RDMLLSKTRE ARETTLPSHS 480 YSKMSQDEEL IEPPVLFRTG SHIVKGIQEQ SLSPRKYPHE QTMLSKESPE KAASLFKQMS 540 IDETGRAAAT ETLHQGEPVY VVDELLPGDS LAGFGSVEEA PGTSIDDVTH PSAQGSRDPS 600 FTSMVQNPFV LGQPGTQEQR QHDSVGGGDK EQDFDFQKAL KNSSRKR* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
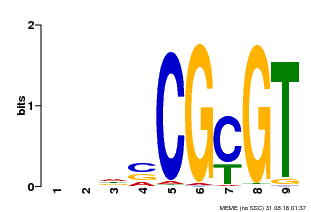
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP5132 | 8 | 9 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 6e-32 | ethylene induced calmodulin binding protein | ||||



