 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010534781.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 450aa MW: 49850.3 Da PI: 6.8381 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 105.5 | 3.1e-33 | 240 | 294 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHerFv+av++L G+ekAtPk++l+lm+v+gLt++hvkSHLQkYRl
XP_010534781.1 240 KPRMRWTPELHERFVQAVHSLEGPEKATPKAVLKLMNVEGLTIYHVKSHLQKYRL 294
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.424 | 237 | 297 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.1E-32 | 238 | 295 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.73E-18 | 238 | 294 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.1E-25 | 240 | 295 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.6E-10 | 242 | 293 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 5.8E-23 | 330 | 377 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 450 aa Download sequence Send to blast |
MNSHSLVSGT ADEGGVGQDC SSSISLVHNF LNFPPETRKA PSPFIRSQSP DSASHDHHPW 60 MKHGSKSTFS RSSSFCTNLY LSSSSSSETR KHLGNSLLPF LPNPSTYNQT ASAVESAKSP 120 AIFSEDIVTV TPPYNGGGSS SSGSLAKDFF NLPKDTCSDG GFHDLGCPND SFSLSEQMEL 180 QFLSDELELA ITDRAETPSL DEIYEMPMAM SNPGTEVSPR QNCVSAVMSH SSPGAATNQK 240 PRMRWTPELH ERFVQAVHSL EGPEKATPKA VLKLMNVEGL TIYHVKSHLQ KYRLAKYIPE 300 KREEKKNSNS EEKKSASSNS ETDGRKKGAI QITEALRMQM EVQKQLHEQL EVQRVLQLRI 360 EEHARYLQRM LEEQQKSGAL ITSSETLSSP TKSKQESSPQ LSSPTKNKAE EENSPKRHRI 420 DNKASESGEE NSPKRPRLDN KASESEERVR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-24 | 240 | 297 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
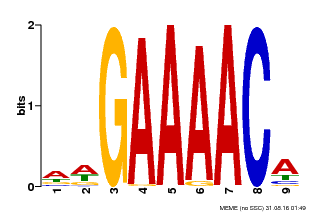
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010534781.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010534781.1 | 0.0 | PREDICTED: myb family transcription factor PHL6-like | ||||
| Refseq | XP_010534782.1 | 0.0 | PREDICTED: myb family transcription factor PHL6-like | ||||
| Refseq | XP_010534783.1 | 0.0 | PREDICTED: myb family transcription factor PHL6-like | ||||
| Refseq | XP_010534785.1 | 0.0 | PREDICTED: myb family transcription factor PHL6-like | ||||
| Refseq | XP_010534786.1 | 0.0 | PREDICTED: myb family transcription factor PHL6-like | ||||
| Swissprot | Q949U2 | 0.0 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | R0HYE1 | 0.0 | R0HYE1_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010534781.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6786 | 27 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-160 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



