 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi06g15690.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 315aa MW: 34095.2 Da PI: 8.9499 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 205.5 | 1.5e-63 | 13 | 156 | 1 | 145 |
DUF822 1 ggsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspessl 94
++s+rkp+w+ErEnn+rRERrRRaiaakiy+GLRaqGny+lpk++DnneVlkALc+eAGw ve+DGttyrkgskp+ a+ agss ++ ss+
Vradi06g15690.1 13 AASRRKPSWRERENNRRRERRRRAIAAKIYSGLRAQGNYNLPKHCDNNEVLKALCAEAGWCVEEDGTTYRKGSKPP-LANGAGSSMRNILFSSS 105
589*************************************************************************.88888888887766666 PP
DUF822 95 q.sslkssalaspvesysaspksssfpspssldsislasaasllpvlsvlsl 145
q s+ ss+ +sp++sy+asp+sssfpsp +ld + + ++l+p++++ sl
Vradi06g15690.1 106 QnPSPFSSSHPSPIPSYQASPSSSSFPSPFRLDVDKD-NVSNLIPYIRNASL 156
66******************************98865.46899999998876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 2.4E-60 | 15 | 154 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0001046 | Molecular Function | core promoter sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 315 aa Download sequence Send to blast |
MTSDGATSAA AAAASRRKPS WRERENNRRR ERRRRAIAAK IYSGLRAQGN YNLPKHCDNN 60 EVLKALCAEA GWCVEEDGTT YRKGSKPPLA NGAGSSMRNI LFSSSQNPSP FSSSHPSPIP 120 SYQASPSSSS FPSPFRLDVD KDNVSNLIPY IRNASLSLPP LRISNSAPVT PPLSSPTSRN 180 PKPIPTWESI AKESMTSFNY PLFAASAPAS PTHRHLYTPA TIPECDESDT STCESSQWVK 240 FQAFAPSASV LPTSPTFNLV KPVVPPGVPD NSIQEMRTSS EEIGVQVMPW VGERIHEVAL 300 DDLELTLGSG KVRS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 6e-30 | 2 | 98 | 357 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 6e-30 | 2 | 98 | 357 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 6e-30 | 2 | 98 | 357 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 6e-30 | 2 | 98 | 357 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 15 | 34 | RRKPSWRERENNRRRERRRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Positive regulator of brassinosteroid (BR) signaling. Transcription factor that activates target gene expression by binding specifically to the DNA sequence 5'-CANNTG-3'(E box) through its N-terminal domain. Can bind individually to the promoter as a homodimer or synergistically as a heterodimer with BIM1, BIM2 or BIM3. The C-terminal domain is probably involved in transcriptional activation (PubMed:12007405, PubMed:15680330, PubMed:18467490, PubMed:19170933). Recruits the transcription elongation factor IWS1 to control BR-regulated gene expression (PubMed:20139304). Forms a trimeric complex with IWS1 and ASHH2/SDG8 to regulate BR-regulated gene expression (PubMed:24838002). Promotes quiescent center (QC) self-renewal by cell divisions in the primary root. Binds to the E-boxes of the BRAVO promoter to repress its expression (PubMed:24981610). {ECO:0000269|PubMed:12007405, ECO:0000269|PubMed:15680330, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20139304, ECO:0000269|PubMed:24838002, ECO:0000269|PubMed:24981610}. | |||||
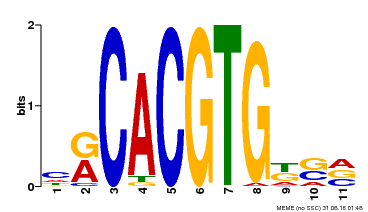
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00073 | ChIP-chip | Transfer from AT1G19350 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi06g15690.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 0.0 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014505170.1 | 0.0 | protein BRASSINAZOLE-RESISTANT 2 | ||||
| Swissprot | Q9LN63 | 1e-117 | BZR2_ARATH; Protein BRASSINAZOLE-RESISTANT 2 | ||||
| TrEMBL | A0A1S3UGL0 | 0.0 | A0A1S3UGL0_VIGRR; protein BRASSINAZOLE-RESISTANT 2 | ||||
| STRING | GLYMA14G08320.1 | 1e-176 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1275 | 34 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19350.3 | 1e-108 | BES1 family protein | ||||



