 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK14402.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 408aa MW: 45390.4 Da PI: 6.7613 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.1 | 7e-32 | 272 | 327 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+ eLH+rFv+a++qLGGs++AtPk+i+elm v+gLt+++vkSHLQkYRl+
PK14402.1 272 KQRRCWSIELHRRFVDAIHQLGGSHVATPKQIRELMRVEGLTNDEVKSHLQKYRLH 327
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.423 | 269 | 329 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.02E-16 | 270 | 330 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-28 | 271 | 330 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-25 | 272 | 327 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.7E-7 | 274 | 325 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 408 aa Download sequence Send to blast |
MELSLNNNSG LAFVPKTISE FLEHVSKIRD GSEKLSKLDD YVKRFEEEKR KIDAFKRELP 60 LCMSLVTDAI ERLKEEEKAI KAMDEFTSFK GNSDENGSTN LVDDSCDKKS WMSSAQLWST 120 NVVDCDSHNK QTSVSELKPR IEEVAVDRHV SEKQPIGVSD YRNRGGAFVP FKAQPSPPSG 180 VIVRTSLKED KEVSLSRVPS LSLMPPTPLP TTSPISEMGG TSGDSNPKSS HSCKTGSASS 240 FLTEQVKLVS KSQLVQHQTQ HQHHQQQQPF RKQRRCWSIE LHRRFVDAIH QLGGSHVATP 300 KQIRELMRVE GLTNDEVKSH LQKYRLHVRK LPNSSTEPDS GLWMSQDQCG DSSHQSASPQ 360 GPLAPSGSNK ALSHTGGNST EVEDEEEDEK SDGHSWRGGN HNKAGEDV |
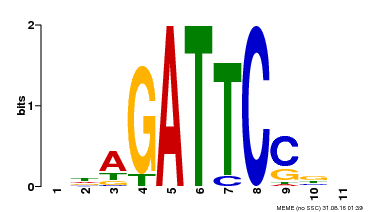
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | 25215497 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015890038.1 | 1e-134 | transcription factor HHO5-like | ||||
| TrEMBL | A0A2P5ADI9 | 0.0 | A0A2P5ADI9_PARAD; GARP transcription factor | ||||
| STRING | XP_010109894.1 | 1e-128 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1208 | 34 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49560.1 | 4e-33 | G2-like family protein | ||||



