 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400044384 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 307aa MW: 35011.8 Da PI: 6.4389 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.9 | 7.2e-18 | 79 | 132 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
PGSC0003DMP400044384 79 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 132
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 132 | 2.2e-42 | 78 | 170 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdye+Lkr++da+k+en++L++++++L++
PGSC0003DMP400044384 78 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYEVLKRQFDAIKAENDALQTQNQKLHA 166
69**************************************************************************************9 PP
HD-ZIP_I/II 90 elke 93
e+++
PGSC0003DMP400044384 167 EIMS 170
9975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.3E-19 | 55 | 135 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.4E-19 | 70 | 136 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 74 | 134 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 77 | 138 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.2E-15 | 79 | 132 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.54E-15 | 79 | 135 | No hit | No description |
| PRINTS | PR00031 | 1.3E-5 | 105 | 114 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 109 | 132 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.3E-5 | 114 | 130 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 8.0E-17 | 134 | 175 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 307 aa Download sequence Send to blast |
MAFFPTNFML QTPHHEDEHQ PSTSLNPILP SCSPQDFHGI ASFLGKRSMS FSGMDGNNAC 60 EENHGEDDLS DDGSQAGEKK RRLNMEQVKT LEKNFELGNK LEPERKMQLA RALGLQPRQI 120 AIWFQNRRAR WKTKQLEKDY EVLKRQFDAI KAENDALQTQ NQKLHAEIMS LKNREQPTES 180 INLNKETEGS CSNRSENSSE IKLDISRTPA IDSPLSNHHP NISSRPFFPP SMIRSNNNNN 240 NNGVVVPHQL FHINSSSSRQ DLKLMDQNTT TTNNNSSVKE ESLSNMFCGI DDQTSFWPWL 300 EQQHFN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 126 | 134 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
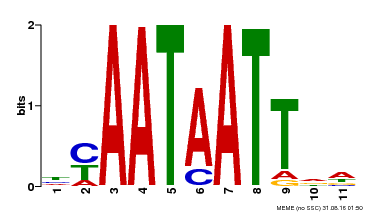
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400044384 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF011556 | 0.0 | AF011556.1 Lycopersicon esculentum jasmonic acid 1 (LEJA1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006363243.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-13-like | ||||
| Swissprot | Q8LC03 | 1e-121 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | M1CEL3 | 0.0 | M1CEL3_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400065805 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA848 | 24 | 98 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-122 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400044384 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



