 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000423596 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 433aa MW: 47701.1 Da PI: 6.1412 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.2 | 1.2e-17 | 227 | 280 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
MDP0000423596 227 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 280
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.7 | 5.8e-42 | 226 | 318 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr++da+k++n++L++++++L++e+++
MDP0000423596 226 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDLLKRQFDAIKADNDALQSQNQKLQAEIMA 318
69**************************************************************************************99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.81E-19 | 218 | 284 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 222 | 282 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 225 | 286 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 5.3E-15 | 227 | 280 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.06E-16 | 227 | 283 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-19 | 229 | 289 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 2.2E-5 | 253 | 262 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 257 | 280 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.2E-5 | 262 | 278 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.3E-16 | 282 | 322 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 433 aa Download sequence Send to blast |
MAGTAIISLI SVTEYSKGIE HVIGGNENIR IASLIVFALL GFPLAMIVSL GAGPWDALFG 60 GGNIPAFVLA SFAALAGGII AARRLPILSS SLNDDVDQGF GIEGVDCGGM ITCHLVIEVV 120 VWQNRSKRVE QAIIKIGGKS LINHQAMTAN GMAFFSTNFM LQTPHDRDDH QPPTSLSPML 180 PSCTPQDFHG VASFLGKRSV SFSGIELGEE AHGEDDLSDD GSQAGEKKRR LNMEQVKTLE 240 KNFELGNKLE PERKMQLARA LGLQPRQIAI WFQNRRARWK TKQLEKDYDL LKRQFDAIKA 300 DNDALQSQNQ KLQAEIMALK NREPAESINL NKDTEGSCSN RSENNSDIKL DISRTPAIDS 360 PPSSHHHQNP TLFSSSIIRP AQLFQNTSRT EAMQCQKIDQ MVKEESLTNM FCGIDDQSAA 420 GFWPWLEQHQ FN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 274 | 282 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.15826 | 0.0 | bud| fruit| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in leaves and flowers. {ECO:0000269|PubMed:16055682}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
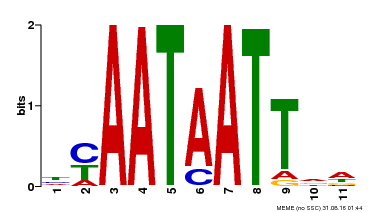
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001280955.1 | 0.0 | homeobox-leucine zipper protein ATHB-13-like | ||||
| Swissprot | Q8LC03 | 1e-119 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | D9ZJ08 | 0.0 | D9ZJ08_MALDO; HD domain class transcription factor | ||||
| STRING | XP_008339924.1 | 0.0 | (Malus domestica) | ||||
| STRING | XP_008352991.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1271 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-121 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000423596 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



