 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os05g47650.1 | ||||||||
| Common Name | LOC4339522, Os05g0549800, P0560C03.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 395aa MW: 41380.9 Da PI: 10.055 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.5 | 1.4e-12 | 67 | 115 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ r +r++lg+f + eAa+a++ a+++++g
LOC_Os05g47650.1 67 SKYKGVVPQP-NGRWGAQIYE------RhQRVWLGTFTGEAEAARAYDVAAQRFRG 115
89****9888.8*********......44**********99*************98 PP
| |||||||
| 2 | B3 | 91.5 | 6.4e-29 | 193 | 306 | 1 | 92 |
EEEE-..-HHHHTT-EE--HHH.HTT.......................---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......................ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvk 70
f+k+ tpsdv+k++rlv+pk++ae+h g +++ l +ed g++W+++++y+++s++yvltkGW++Fvk
LOC_Os05g47650.1 193 FDKTVTPSDVGKLNRLVIPKQHAEKHfplqlppptttssvaaaadaaagG-GDCKGVLLNFEDAAGKVWKFRYSYWNSSQSYVLTKGWSRFVK 284
89***************************888888887777777777643.334999************************************ PP
HHT--TT-EEEEEE-SSSEE.. CS
B3 71 angLkegDfvvFkldgrsefel 92
++gL +gD v F++ ++++l
LOC_Os05g47650.1 285 EKGLHAGDAVGFYRAAGKNAQL 306
*************765555555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.7E-16 | 67 | 123 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.7E-8 | 67 | 115 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.50E-23 | 67 | 123 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.5E-19 | 68 | 123 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.2E-26 | 68 | 129 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.862 | 68 | 123 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 1.4E-39 | 184 | 320 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.50E-26 | 191 | 298 | No hit | No description |
| SuperFamily | SSF101936 | 2.62E-28 | 191 | 311 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.097 | 193 | 314 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.8E-26 | 193 | 308 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 9.2E-21 | 193 | 313 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0020039 | anatomy | leaf lamina | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 395 aa Download sequence Send to blast |
MDSTSCLLDD ASSGASTGKK AAAAAASKAL QRVGSGASAV MDAAEPGAEA DSGGERRGGG 60 GGKLPSSKYK GVVPQPNGRW GAQIYERHQR VWLGTFTGEA EAARAYDVAA QRFRGRDAVT 120 NFRPLAESDP EAAVELRFLA SRSKAEVVDM LRKHTYLEEL TQNKRAFAAI SPPPPKHPAS 180 SPTSSSAARE HLFDKTVTPS DVGKLNRLVI PKQHAEKHFP LQLPPPTTTS SVAAAADAAA 240 GGGDCKGVLL NFEDAAGKVW KFRYSYWNSS QSYVLTKGWS RFVKEKGLHA GDAVGFYRAA 300 GKNAQLFIDC KVRAKPTTAA AAAAFLSAVA AAAAPPPAVK AIRLFGVDLL TAAAPELQDA 360 GGAAMTKSKR AMDAMAESQA HVVFKKQCIE LALT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 3e-49 | 190 | 314 | 11 | 118 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.9064 | 1e-133 | callus| leaf| root| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | Q6L4H4 | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
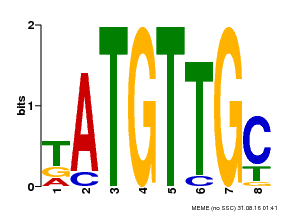
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os05g47650.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os05g47650 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC130725 | 0.0 | AC130725.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0494H05, complete sequence. | |||
| GenBank | AC135925 | 0.0 | AC135925.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0560C03, complete sequence. | |||
| GenBank | AC136492 | 0.0 | AC136492.3 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0554F08, complete sequence. | |||
| GenBank | AP014961 | 0.0 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015637817.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| Swissprot | Q6L4H4 | 0.0 | Y5498_ORYSJ; AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| TrEMBL | A0A0E0HHS8 | 0.0 | A0A0E0HHS8_ORYNI; Uncharacterized protein | ||||
| STRING | ONIVA05G26000.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 | Representative plant | OGRP365 | 16 | 108 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 2e-95 | RAV family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os05g47650.1 |
| Entrez Gene | 4339522 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



