 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os03g14270.1 | ||||||||
| Common Name | LOC4332242, Os03g0246900, OsJ_10128, OSNPB_030246900 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 276aa MW: 28252.8 Da PI: 8.1882 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 130.2 | 8.9e-41 | 37 | 137 | 1 | 100 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeq.pkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqqleqlka 92
+C aCk+lrrkC+++C++apyf +eq +++fa+vhk+FGasnv+kll ++p ++r+da+ +++yeA+ar+rdPvyG+v++i++lqqq+ +l+a
LOC_Os03g14270.1 37 PCGACKFLRRKCVSGCIFAPYFDSEQgAAHFAAVHKVFGASNVSKLLLQIPAHKRPDAVVTICYEAQARLRDPVYGCVAHIFALQQQVVNLQA 129
7***********************9989***************************************************************** PP
DUF260 93 elallkee 100
el +l+++
LOC_Os03g14270.1 130 ELTYLQAH 137
***99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF038127 | 2.8E-134 | 1 | 275 | IPR017394 | LOB domain-containing protein 18/20 |
| PROSITE profile | PS50891 | 23.348 | 36 | 138 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 5.2E-40 | 37 | 135 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010199 | Biological Process | organ boundary specification between lateral organs and the meristem | ||||
| GO:0010311 | Biological Process | lateral root formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009049 | anatomy | inflorescence | ||||
| PO:0001083 | developmental stage | inflorescence development stage | ||||
| PO:0007006 | developmental stage | IL.00 inflorescence just visible stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 276 aa Download sequence Send to blast |
MSSGGGASSA LGGGGGGGGS GGGGGGPSGG GGGGGGPCGA CKFLRRKCVS GCIFAPYFDS 60 EQGAAHFAAV HKVFGASNVS KLLLQIPAHK RPDAVVTICY EAQARLRDPV YGCVAHIFAL 120 QQQVVNLQAE LTYLQAHLAT LELPSPPPLM PAPPQMPMPA PFSISDLPSS TSVPTTVDLS 180 ALFDPPPQPQ WASPLQQQQH HHHQHHHHQQ QQHQLRQPSY ATLARAPSGM TAAAESSGGG 240 GGGGGGDLQA LARELLDRHR SAVKLEQPPP PHSRS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 1e-37 | 33 | 131 | 7 | 104 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 1e-37 | 33 | 131 | 7 | 104 | LOB family transfactor Ramosa2.1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.83177 | 0.0 | panicle | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 115451890 | 0.0 | ||||
| Expression Atlas | Q10P52 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: During lateral root formation, expressed in the lateral root primordia, and the developing, emerged, and mature lateral roots. {ECO:0000269|PubMed:19717544}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and flowers (PubMed:12068116). Expressed in vascular tissues of hypocotyls, leaves, roots, developing floral organs and siliques (PubMed:19088331). {ECO:0000269|PubMed:12068116, ECO:0000269|PubMed:19088331}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and flowers (PubMed:12068116, PubMed:24484953). Expressed in vascular tissues of hypocotyls, leaves, roots, developing floral organs and siliques (PubMed:19088331). {ECO:0000269|PubMed:12068116, ECO:0000269|PubMed:19088331, ECO:0000269|PubMed:24484953}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the positive regulation of tracheary element (TE) differentiation. Involved in a positive feedback loop that maintains or promotes NAC030/VND7 expression that regulates TE differentiation-related genes. {ECO:0000269|PubMed:19088331}. | |||||
| UniProt | Involved in the positive regulation of tracheary element (TE) differentiation. Involved in a positive feedback loop that maintains or promotes NAC030/VND7 expression that regulates TE differentiation-related genes (PubMed:19088331). Functions in the initiation and emergence of lateral roots, in conjunction with LBD16, downstream of ARF7 and ARF19 (PubMed:19717544, PubMed:23749813). Transcriptional activator that directly regulates EXPA14, a gene encoding a cell wall-loosening factor that promotes lateral root emergence. Activates EXPA14 by directly binding to a specific region of its promoter (PubMed:22974309). Transcriptional activator that directly regulates EXPA17, a gene encoding a cell wall-loosening factor that promotes lateral root emergence (PubMed:23872272). Acts downstream of the auxin influx carriers AUX1 and LAX1 in the regulation of lateral root initiation and development (PubMed:26059335). {ECO:0000269|PubMed:19088331, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:22974309, ECO:0000269|PubMed:23749813, ECO:0000269|PubMed:23872272, ECO:0000269|PubMed:26059335}. | |||||
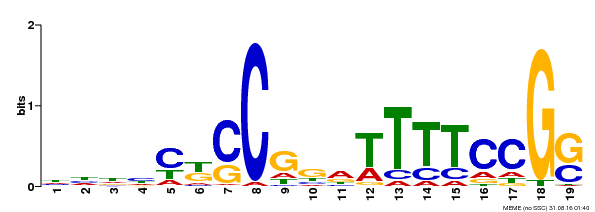
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00314 | DAP | Transfer from AT2G45420 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os03g14270.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:15659631, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:23749813}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os03g14270 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC118672 | 0.0 | AC118672.3 Genomic sequence for Oryza sativa, Nipponbare strain, clone OSJNBb0078P24, from chromosome 3, complete sequence. | |||
| GenBank | AP014959 | 0.0 | AP014959.1 Oryza sativa Japonica Group DNA, chromosome 3, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012611 | 0.0 | CP012611.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 3 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015629413.1 | 0.0 | LOB domain-containing protein 30 | ||||
| Swissprot | O22131 | 8e-70 | LBD18_ARATH; LOB domain-containing protein 18 | ||||
| Swissprot | O81323 | 3e-70 | LBD30_ARATH; LOB domain-containing protein 30 | ||||
| TrEMBL | Q10P52 | 0.0 | Q10P52_ORYSJ; LOB domain protein 18, putative | ||||
| STRING | OS03T0246900-00 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3346 | 37 | 82 | Representative plant | OGRP405 | 15 | 100 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00220.1 | 7e-68 | LBD family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os03g14270.1 |
| Entrez Gene | 4332242 |



