 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN49105.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 409aa MW: 43601.4 Da PI: 6.502 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 12.6 | 0.00026 | 249 | 270 | 35 | 55 |
TS-.STCHHHHHHHHHHHHHHH CS
HLH 35 skK.lsKaeiLekAveYIksLq 55
sk +Ka++L +++eY+k+Lq
KHN49105.1 249 SKPmTDKASMLDEVIEYLKQLQ 270
555589***************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.280.10 | 5.9E-6 | 244 | 280 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 5.52E-5 | 252 | 285 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 0.00132 | 253 | 274 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009567 | Biological Process | double fertilization forming a zygote and endosperm | ||||
| GO:0009506 | Cellular Component | plasmodesma | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 409 aa Download sequence Send to blast |
MSQCVPSWDV EDNPPPSRVS LRSNSNSTAP DVPMLDYEVA ELTWENGQLS MHGLGLPRVP 60 VKPPTAATNK YTWEKPRGSG TLESIVNQAT SFSHQEKPRP LNGDSGGGGG VYGNFMVPWF 120 DPHAAATTTT TTTNTMTMDA LVPCSNREQG KKKGMEPGPG TCMVGCSTRV GSCCGGKGAK 180 GHEASGRDQS VSGSATFGRD SKHVTLDTCD REFGVAFTST SINSLENTSY AKHCTKTTTI 240 EEHDSVSHSK PMTDKASMLD EVIEYLKQLQ AQVQMMNRIN MSSMMLPLTM QQQLQMSMMS 300 PMGMGLGMGM GMGMGMGMDM NSMNRANIPG IPPVLHPSAF MPMAASWDAA AAAAAGGGDR 360 LQGTPASVMP DPLSTIFGCQ SQPMTMDAYS RLAAMYQQLH QPPTSGSKN |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00034 | PBM | Transfer from AT4G00050 | Download |
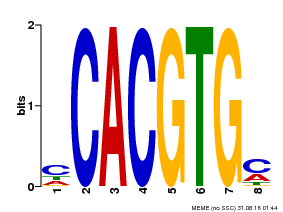
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN49105.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235052 | 0.0 | AC235052.1 Glycine max strain Williams 82 clone GM_WBb0119P14, complete sequence. | |||
| GenBank | AC235252 | 0.0 | AC235252.1 Glycine max strain Williams 82 clone GM_WBb0036F08, complete sequence. | |||
| GenBank | AC235273 | 0.0 | AC235273.1 Glycine max strain Williams 82 clone GM_WBb0046P21, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028187769.1 | 0.0 | transcription factor UNE10-like | ||||
| TrEMBL | A0A0B2SW75 | 0.0 | A0A0B2SW75_GLYSO; Transcription factor UNE10 | ||||
| STRING | GLYMA11G17120.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5676 | 33 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00050.1 | 3e-60 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



