|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Glyma.03G255000.2.p |
| Common Name | GLYMA_03G255000 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
| Family |
bZIP |
| Protein Properties |
Length: 425aa MW: 45528.4 Da PI: 8.2266 |
| Description |
bZIP family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Glyma.03G255000.2.p | genome | JGI | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | bZIP_1 | 70.6 | 2.4e-22 | 279 | 341 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkrerrkq+NRe+ArrsR+RK+ae+eeL++kv++L aeN +Lk+e+++l++ ++k++ e+
Glyma.03G255000.2.p 279 ERELKRERRKQSNRESARRSRLRKQAETEELARKVESLNAENATLKSEINRLTESSEKMRVEN 341
89*********************************************************9987 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated |
| GO:0009737 | Biological Process | response to abscisic acid |
| GO:0005829 | Cellular Component | cytosol |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0043565 | Molecular Function | sequence-specific DNA binding |
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 425 aa
Download sequence Send
to blast |
MGNSEEEKST KTEKPSSPVT MDQANQTNQT NIHVYPDWAA MQAYYGPRVT MPPYYNSAVA 60
SGHAPHPYMW GPPQPMMPPY GPPYAAIYPH GGVYTHPAVP IGPLTHSQGV PSSPAAGTPL 120
SIETPPKSSG NTDQGLMKKL KEFDGLAMSI GNGHAESAER GGENRLSQSV DTEGSSDGSD 180
GNTSGANQSR RKRSREGTPT TDGEGKTEIQ GSPISKETAA SNKMLGVVPA SVAGTIVGHV 240
VSSGMTTALE LRNPSSVHSK TSAPQPCPVL PAEAWVQNER ELKRERRKQS NRESARRSRL 300
RKQAETEELA RKVESLNAEN ATLKSEINRL TESSEKMRVE NATLRGKLKN AQLGQTQEIT 360
LKIIDSQRAT PVSTENLLSR VNNSGSNDRT VEDENGFCEN KPNSGAKLHQ LLDTSPRADA 420
VAAG*
|
| Nucleic Localization
Signal ? help
Back to Top |
 |
| No. |
Start |
End |
Sequence |
| 1 | 295 | 301 | RRSRLRK |
| 2 | 295 | 302 | RRSRLRKQ |
| Functional Description ? help
Back to Top |
| Source |
Description |
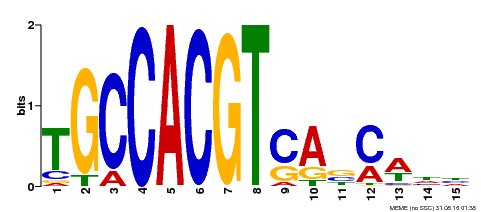
| UniProt | Binds to the G-box-like motif (5'-ACGTGGC-3') of the chalcone synthase (CHS) gene promoter. G-box and G-box-like motifs are defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: By light. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | KT031145 | 0.0 | KT031145.1 Glycine max clone HN_CCL_210 BZIP transcription factor (Glyma19g44190.2) mRNA, partial cds. |