 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10013698m | ||||||||
| Common Name | CARUB_v10013698mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 447aa MW: 49374.9 Da PI: 6.5743 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100 | 1.6e-31 | 236 | 290 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHe+Fv+ v +L G+ekAtPk++l+lm+v+gLt++hvkSHLQkYRl
Carubv10013698m 236 KPRMRWTPELHESFVNSVIKLEGPEKATPKAVLKLMNVEGLTIYHVKSHLQKYRL 290
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-29 | 234 | 291 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.5E-16 | 235 | 290 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-23 | 236 | 291 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.7E-7 | 238 | 289 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.5E-22 | 326 | 373 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 447 aa Download sequence Send to blast |
MNRHSLVSVA PDKCNEEVGQ SCSSSLSPVH NFLNVQPETR KSPFIRSQSP DSPGQLWPKH 60 SSQSTFSRSS TFCTNLYLSS SSTSETQKHL GNCLPFLPDP SSYSHSASGV ESARSPSIFS 120 EDLGNQYDGN NSGSLVKDFL NLSGDVCSDG GFHDFGCSND SYCLSDQMEL QFLSDELELA 180 ISDRAETPRL DEIYETPLAS VNPVTRLSPS QSCVAGAVST DVVSSHPSPG SAANHKPRMR 240 WTPELHESFV NSVIKLEGPE KATPKAVLKL MNVEGLTIYH VKSHLQKYRL AKYMPEKKEG 300 KKNDNSEEKK LAFSNSEADG KRKGAIQLTE ALRMQMEVQK QLHEQLEVQR VLQLRIEEHA 360 KYLEKMLEEQ RKTGRLISSS SPSQTVLSPS DESIENSENL SKTKASSPQP SSSTKNKASE 420 TEGNMCESPQ KRRRLENKAE SEDPKR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-23 | 236 | 293 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 430 | 434 | KRRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
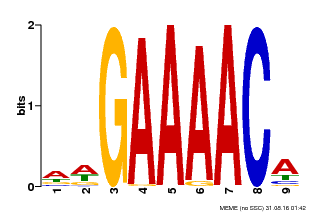
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10013698m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY050889 | 0.0 | AY050889.1 Arabidopsis thaliana unknown protein (At3g13040) mRNA, complete cds. | |||
| GenBank | AY096674 | 0.0 | AY096674.1 Arabidopsis thaliana unknown protein (At3g13040) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006297672.1 | 0.0 | myb family transcription factor PHL6 | ||||
| Swissprot | Q949U2 | 0.0 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | R0HYE1 | 0.0 | R0HYE1_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006297672.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6786 | 27 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 0.0 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10013698m |
| Entrez Gene | 17892662 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



