 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG003168.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 809aa MW: 88985.9 Da PI: 6.6211 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
CCG003168.2 17 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 169.4 | 2.5e-53 | 161 | 368 | 2 | 204 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEEEX CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmvae 97
+aee+++e+++ka+ ++ Wv+++ +++g+++ +++ s++++g +ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l +++
CCG003168.2 161 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSSGIIAISHGCTGVGARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTIELLYMQ 257
789*******************************************************.8888888888****************9999********* PP
XTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHH CS
START 98 lqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglae 191
l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+l++p+++g+s +++v+h+dl+ ++++++lr+l++s+++
CCG003168.2 258 LYAPTTLAPgRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPSmppVQHFVRAEMLPSGYLVRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLYESSTVL 355
****************************************9988899*************************************************** PP
HHHHHHHHTXXXX CS
START 192 gaktwvatlqrqc 204
++kt++ +l++++
CCG003168.2 356 AQKTTMVALRQLR 368
******9999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.273 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.41E-16 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.24E-16 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 5.6E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.09E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 24.897 | 151 | 366 | IPR002913 | START domain |
| CDD | cd08875 | 2.65E-76 | 155 | 369 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 4.5E-22 | 160 | 367 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 8.93E-36 | 160 | 372 | No hit | No description |
| SMART | SM00234 | 9.3E-36 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 8.0E-51 | 161 | 368 | IPR002913 | START domain |
| Pfam | PF08670 | 6.4E-29 | 695 | 792 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 809 aa Download sequence Send to blast |
MAMSCKDGKQ PIMDNGKYVR YTPEQVEALE RLYHDCPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQH 120 THNTPLATKD TSCESVVTSG QHHLKPQHPP RDASPAGLLS IAEETLTEFL SKATGTAVEW 180 VQMPGMKPGP DSSGIIAISH GCTGVGARAC GLVGLEPTRV AEILKDRPSW FRDCRAVDVL 240 NVLPTANGGT IELLYMQLYA PTTLAPGRDF WLLRYTSVLE DGSLVVCERS LKNTQNGPSM 300 PPVQHFVRAE MLPSGYLVRP CEGGGSIIHI VDHMDLEPWS VPEVLRPLYE SSTVLAQKTT 360 MVALRQLRQM AQEASQSNVT NWGRRPAALR ALSQRLSRGF NEALNGFSDE GWSMIGNDGM 420 DDVTILVNSS PDKLMGSNLS FTNGFPAVSS AVLCAKASML LQNVPPAILL RFLREHRSEW 480 ADNNIDAYAA AAVKVGPFSL QGSRVGSFGG QVILPLAHTI EHEEFLEVIK LEGVGHSPED 540 PIMPRDVFLL QLCCGMDENA VGACAELIFA PIDATFADDA PLLPSGFRII PLDSGKEASS 600 PNRTLDLAAA LEVGPAGNRA SSDHSANSGC TRSVMTIAFE FAFESHMQEH VASMTRQYIR 660 SIISSVQRVA LALSPSHLGS QAGLRSPLGT PEAQTLARWI CQSYRSYLGV ELLKSNGEGS 720 ESILKTLWHH SDAIMCCSLK VLPIFTFANQ AGLDMLETTL VALQDITLEK IFDDHGRKTL 780 CSEFPQIMQQ VFPSLLNLSE GKMNPVFPA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
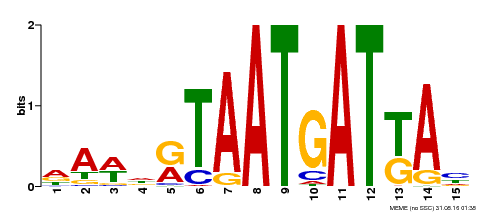
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF534716 | 0.0 | KF534716.1 Populus alba x Populus glandulosa HD-ZIPIII mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011013041.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | V5MYH9 | 0.0 | V5MYH9_9ROSI; HD-ZIPIII | ||||
| STRING | POPTR_0003s04860.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.3 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



