 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G08500.1 | ||||||||
| Common Name | AtMYB83, MYB83, T8G24.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 343aa MW: 38418.8 Da PI: 6.0972 | ||||||||
| Description | myb domain protein 83 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.9 | 3e-15 | 32 | 79 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W+++Ede+l+++ G g+W+ Iar g+ R++k+c++rw +yl
AT3G08500.1 32 KGLWSPDEDEKLIRYMLTNGQGCWSDIARNAGLLRCGKSCRLRWINYL 79
678*****************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.1 | 2.7e-15 | 85 | 129 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg+++++E++l+ +++ lG++ W+ Ia +++ gRt++++k++w++
AT3G08500.1 85 RGSFSPQEEDLIFHLHSILGNR-WSQIATRLP-GRTDNEIKNFWNST 129
899*******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-22 | 26 | 82 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.309 | 27 | 79 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.15E-27 | 29 | 126 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-12 | 31 | 81 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.7E-14 | 32 | 79 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.98E-10 | 35 | 79 | No hit | No description |
| PROSITE profile | PS51294 | 24.984 | 80 | 134 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-25 | 83 | 135 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.4E-15 | 84 | 132 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-14 | 85 | 129 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.76E-11 | 87 | 130 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000981 | Molecular Function | RNA polymerase II transcription factor activity, sequence-specific DNA binding | ||||
| GO:0001135 | Molecular Function | transcription factor activity, RNA polymerase II transcription factor recruiting | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 343 aa Download sequence Send to blast |
MMMRKPDITT IRDKGKPNHA CGGNNNKPKL RKGLWSPDED EKLIRYMLTN GQGCWSDIAR 60 NAGLLRCGKS CRLRWINYLR PDLKRGSFSP QEEDLIFHLH SILGNRWSQI ATRLPGRTDN 120 EIKNFWNSTL KKRLKNNSNN NTSSGSSPNN SNSNSLDPRD QHVDMGGNST SLMDDYHHDE 180 NMMTVGNTMR MDSSSPFNVG PMVNSVGLNQ LYDPLMISVP DNGYHQMGNT VNVFSVNGLG 240 DYGNTILDPI SKRVSVEGDD WFIPPSENTN VIACSTSNNL NLQALDPCFN SKNLCHSESF 300 KVGNVLGIEN GSWEIENPKI GDWDLDGLID NNSSFPFLDF QVD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 9e-29 | 28 | 134 | 3 | 108 | B-MYB |
| 1h8a_C | 9e-29 | 28 | 134 | 23 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 256155_at | 0.0 | ||||
| Expression Atlas | AT3G08500 | - | ||||
| AtGenExpress | AT3G08500 | - | ||||
| ATTED-II | AT3G08500 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed specifically in fiber and vessel cells that are undergoing secondary wall thickening in floral stems. Expressed in vessels but not in xylary fibers in the developing secondary xylem of roots. {ECO:0000269|PubMed:19808805}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a putative R2R3-type MYB transcription factor (MYB83). | |||||
| UniProt | Transcription factor that acts as molecular switch in the NAC012/SND1-mediated transcriptional network regulating secondary wall biosynthesis. Is directly activated by NAC012/SND1 and its close homologs, including NAC043/NST1, NAC066/NST2, NAC101/VND6 and NAC030/VND7. Is required for functional expression of a number of secondary wall-associated transcription factors and secondary wall biosynthetic genes involved in cellulose, xylan and lignin synthesis. Functions redundantly with MYB46 in the transcriptional regulatory cascade leading to secondary wall formation in fibers and vessels (PubMed:19808805). Transcription activator that binds to the DNA consensus sequence 5'-ACC[AT]A[AC][TC]-3', designated as the secondary wall MYB-responsive element (SMRE). Regulates directly numerous transcription factors and a number of genes involved in secondary wall biosynthesis that contain SMRE elements in their promoters (PubMed:22197883). {ECO:0000269|PubMed:19808805, ECO:0000269|PubMed:22197883}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
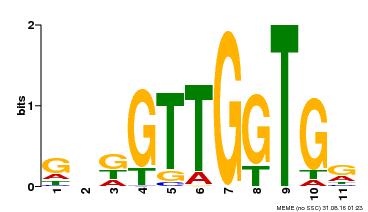
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00336 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G08500.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G08500 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF371974 | 0.0 | AF371974.1 Arabidopsis thaliana putative transcription factor MYB83 (MYB83) mRNA, complete cds. | |||
| GenBank | BT026506 | 0.0 | BT026506.1 Arabidopsis thaliana At3g08500 mRNA, complete cds. | |||
| GenBank | DQ446645 | 0.0 | DQ446645.1 Arabidopsis thaliana clone pENTR221-At3g08500 myb family transcription factor (At3g08500) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_187463.1 | 0.0 | myb domain protein 83 | ||||
| Swissprot | Q9C6U1 | 0.0 | MYB83_ARATH; Transcription factor MYB83 | ||||
| TrEMBL | A0A178V8Z2 | 0.0 | A0A178V8Z2_ARATH; MYB83 | ||||
| STRING | AT3G08500.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7077 | 26 | 42 | Representative plant | OGRP5 | 17 | 1784 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G08500.1 |
| Entrez Gene | 819997 |
| iHOP | AT3G08500 |
| wikigenes | AT3G08500 |



