 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678296432 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 545aa MW: 58860.5 Da PI: 6.3376 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.8 | 3.4e-15 | 41 | 88 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed l d+vk+ G g+W+++ ++ g+ R++k+c++rw ++l
678296432 41 KGPWTSAEDAVLMDYVKKNGEGNWNAVQKHTGLARCGKSCRLRWANHL 88
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 23.8 | 1.1e-07 | 94 | 126 | 1 | 35 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgR 35
+g++T eE+++++++++++G++ W++ a++ +
678296432 94 KGAFTLEEEKIIIELHAKMGNK-WARMAAHAS-EK 126
799*******************.*****9987.55 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 22.33 | 36 | 92 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.8E-11 | 40 | 90 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-13 | 41 | 88 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.7E-23 | 42 | 95 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.59E-24 | 42 | 118 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.82E-9 | 43 | 88 | No hit | No description |
| PROSITE profile | PS51294 | 8.451 | 93 | 174 | IPR017930 | Myb domain |
| SMART | SM00717 | 4.2E-9 | 93 | 172 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-6 | 94 | 127 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.62E-6 | 96 | 168 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-20 | 96 | 121 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-20 | 153 | 172 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 545 aa Download sequence Send to blast |
MGQNSASESE DGAYSANSPL SCNEGDSIGG KSSDNGMILK KGPWTSAEDA VLMDYVKKNG 60 EGNWNAVQKH TGLARCGKSC RLRWANHLRP NLKKGAFTLE EEKIIIELHA KMGNKWARMA 120 AHASEKSDSG FHKGGLFKLY FMEPSFICLK ILSLPGRTDN EIKNYWNTRI KRHQRAGLPI 180 YPPNLYLPAQ SKSHQPLSSP SNCSGDFACH DDTLLQFNGH TKSDAMSGLF SIFQQKPEYG 240 ISPRHSCDDV FLSPHGYTIA QGPDKPILGH ESSDASEPPL FGTTYGAINC HSQSFSMKFS 300 GDLSFDRDTH CVSSIFPSAS KHPIDAQKLE LPSLQYQETN LGFWDPKSKP SDAVTHSPVS 360 GGLSWNCPYS GSSGLLEDLI YGAKALSKPE SHSACWATSC DVSAGDATGD SALNACVTGK 420 VNFNPISTLD NAGGSVPNGR TPTSTMGSSL EDIIVHHLKS ETADERWTLV DEMKNGTHSL 480 DYSRPDTLLE SCGNLQNSLL PPENCPIGSS EASLGDDSKN RSSTSGLGSC PWNNMPAACQ 540 ISEIP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-26 | 39 | 172 | 5 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
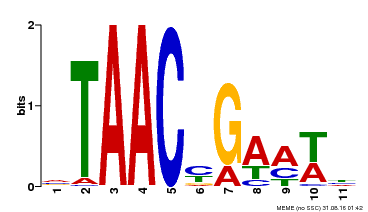
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678296432 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011091869.1 | 1e-158 | transcription factor GAMYB | ||||
| Refseq | XP_011091870.1 | 1e-158 | transcription factor GAMYB | ||||
| Refseq | XP_011091873.1 | 1e-158 | transcription factor GAMYB | ||||
| Refseq | XP_020552724.1 | 1e-158 | transcription factor GAMYB | ||||
| TrEMBL | A0A059PRV5 | 1e-159 | A0A059PRV5_SALMI; MYB-related transcription factor | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA14059 | 7 | 13 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 3e-81 | myb domain protein 33 | ||||



